- java.lang.Object
-
- de.jstacs.clustering.distances.DistanceMetric<StatisticalModel>
-
- de.jstacs.clustering.distances.SequenceScoreDistance
-
- Direct Known Subclasses:
- RandomSequenceScoreDistance
public class SequenceScoreDistance extends DistanceMetric<StatisticalModel>
Class for a distance metric betweenStatisticalModels based on the correlation of score profiles on De Bruijn sequences. For twoStatisticalModels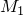 and
and 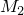 , we compute the score profiles
, we compute the score profiles
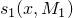 and
and 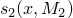 on a De Bruijn sequence
on a De Bruijn sequence 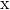 of length
of length 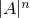 .
The distance is then defined based on the Pearson correlation as
.
The distance is then defined based on the Pearson correlation as 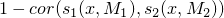 between these score profiles,
maximizing over suitable shifts of the score profiles and both strand orientations.
between these score profiles,
maximizing over suitable shifts of the score profiles and both strand orientations.- Author:
- Jan Grau
-
-
Field Summary
Fields Modifier and Type Field and Description protected booleanexpif exponential scores should be usedprotected CyclicSequenceAdaptor[]seqsThe De Bruijn sequences
-
Constructor Summary
Constructors Modifier Constructor and Description protectedSequenceScoreDistance(CyclicSequenceAdaptor[] seqs, boolean exp)Creates a new distance for a given set of sequences.SequenceScoreDistance(DiscreteAlphabet alphabet, int n, boolean exp)Creates a new distance.
-
Method Summary
All Methods Instance Methods Concrete Methods Modifier and Type Method and Description doublegetDistance(double[][] profiles1, double[][] profiles1Rc, double[][] profiles2, int maxShift)Returns the distance between the two score profiles.doublegetDistance(double[][] profiles1, double[][] profiles1Rc, StatisticalModel o2, int motif1Length)Returns the distance between a score profile and a model.doublegetDistance(StatisticalModel o1, StatisticalModel o2)Returns the distance according to the metric of the two supplied objects.double[][]getPairwiseDistanceMatrix(int numThreads, StatisticalModel... objects)Multi-threaded computation of the pairwise distance matrix.double[][]getProfile(StatisticalModel o, boolean rc)Returns the score profile for the model.-
Methods inherited from class de.jstacs.clustering.distances.DistanceMetric
getPairwiseDistanceMatrix
-
-
-
-
Field Detail
-
seqs
protected CyclicSequenceAdaptor[] seqs
The De Bruijn sequences
-
exp
protected boolean exp
if exponential scores should be used
-
-
Constructor Detail
-
SequenceScoreDistance
public SequenceScoreDistance(DiscreteAlphabet alphabet, int n, boolean exp) throws WrongAlphabetException, WrongSequenceTypeException
Creates a new distance.- Parameters:
alphabet- the alphabet of the models that may be comparedn- the length of n-mers represented in the De Bruijn sequenceexp- if exponential scores should be used- Throws:
WrongAlphabetException- if the sequence of this alphabet could not be createdWrongSequenceTypeException- if the sequence could not be created
-
SequenceScoreDistance
protected SequenceScoreDistance(CyclicSequenceAdaptor[] seqs, boolean exp)
Creates a new distance for a given set of sequences.- Parameters:
seqs- the sequencesexp- if exponential scores should be used
-
-
Method Detail
-
getProfile
public double[][] getProfile(StatisticalModel o, boolean rc) throws Exception
Returns the score profile for the model.- Parameters:
o- the moderc- if the reverse complement should be considered- Returns:
- the score profile
- Throws:
Exception- if the score could not be computed
-
getDistance
public double getDistance(double[][] profiles1, double[][] profiles1Rc, double[][] profiles2, int maxShift) throws ExceptionReturns the distance between the two score profiles.- Parameters:
profiles1- the first profileprofiles1Rc- the reverse complementary version of the first profileprofiles2- the second profilemaxShift- the maximum allowed shift between the profiles- Returns:
- the distance
- Throws:
Exception- if the distance could not be computed
-
getDistance
public double getDistance(double[][] profiles1, double[][] profiles1Rc, StatisticalModel o2, int motif1Length) throws ExceptionReturns the distance between a score profile and a model.- Parameters:
profiles1- the first profileprofiles1Rc- the reverse complementary version of the first profileo2- the modelmotif1Length- the length of the motif used to compute the first profile- Returns:
- the distance
- Throws:
Exception- if the distance could not be computed
-
getDistance
public double getDistance(StatisticalModel o1, StatisticalModel o2) throws Exception
Description copied from class:DistanceMetricReturns the distance according to the metric of the two supplied objects.- Specified by:
getDistancein classDistanceMetric<StatisticalModel>- Parameters:
o1- the first objecto2- the second object- Returns:
- the distance
- Throws:
Exception- if the distance could not be computed
-
getPairwiseDistanceMatrix
public double[][] getPairwiseDistanceMatrix(int numThreads, StatisticalModel... objects) throws ExceptionMulti-threaded computation of the pairwise distance matrix.- Parameters:
numThreads- the number of threadsobjects- the models- Returns:
- the distance matrix
- Throws:
Exception- if the distance could not be computed
-
-