AnnoTALE
Transcription activator-like effectors (TALEs) are virulence factors of plant-pathogenic Xanthomonas spp. that function as gene activators inside plant host cells.
AnnoTALE is a suite of applications for identifying and analysing TALEs in Xanthomonas genomes, for clustering TALEs into classes by their RVD sequences, for assigning novel TALEs to existing classes, for proposing TALE names using a unified nomenclature, and for predicting targets of individual TALEs and TALE classes.
AnnoTALE is available as a JavaFX-based stand-alone application with graphical user interface for interactive analysis sessions. In addition, we provide a command line application that may be integrated into other pipelines. Both use identical code for the actual analysis, ensuring consistent results between both versions.
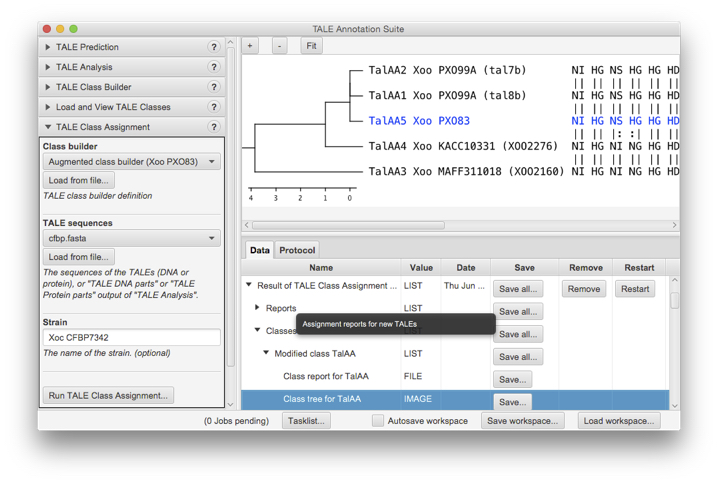
AnnoTALE with GUI
AnnoTALE is based on the very recent implementation of JavaFX in Java 8.
We provide AnnoTALE as a runnable JAR file for those with a current version of Java 8 (at least update 45) on their machine.
For user's convenience, we also provide pre-packaged versions of AnnoTALE, which also include Java in the required version, for Mac OS X and Windows. Each of these versions is available two version with different memory requirements (2GB and 6GB). As long as the main memory (RAM) of your machine is sufficient, we recommend to use the 6GB version of AnnoTALE.
Download
AnnoTALE is distributed in the hope that it will be useful, but without any warranty; without even the implied warranty of merchantability or fitness for a particular purpose.
- Runnable Jar (requires Java 8, update 45 or greater)
- Mac-DMG of AnnoTALE including Java: 2GB version, 6GB version
- Windows installer of AnnoTALE including Java: 1GB version, 2GB version, 6GB version
User Guide
We provide an AnnoTALE User Guide in PDF format, including a detailed description of all AnnoTALE tools and installation instructions.
AnnoTALE command line application
The AnnoTALE command line application is available as a runnable Jar. For running the program and a quick help, type
java -jar AnnoTALEcli-1.0.jar
For larger analyes, it might be necessary to increase the memory allocated by the JavaVM using the -Xms and -Xmx parameters, for instance
java -Xms512M -Xmx6G -jar AnnoTALEcli-1.0.jar
There is no separate User Guide for the AnnoTALE command line application, but the User Guide for the GUI version describes all AnnoTALE tools, their parameters and outputs, and those of the CLI version are identical.
You obtain a list of all AnnoTALE tools by calling
java -jar AnnoTALEcli-1.0.jar
Output:
Available tools: predict - TALE Prediction analyze - TALE Analysis build - TALE Class Builder loadAndView - Load and View TALE Classes assign - TALE Class Assignment rename - Rename TALEs in File targets - Predict and Intersect Targets Syntax: java -jar AnnoTALEcli-1.0.jar <toolname> [<parameter=value> ...] Further info about the tools is given with java -jar AnnoTALEcli-1.0.jar <toolname> info Tool parameters are listed with java -jar AnnoTALEcli-1.0.jar <toolname>
You get a list of input parameters by calling AnnoTALEcli-1.0.jar with the corresponding tool name, e.g.,
java -jar AnnoTALEcli-1.0.jar predict
Output:
At least one parameter has not been set (correctly): Parameters of tool "TALE Prediction" (predict): g - Genome (The input Xanthomonas genome in FastA or Genbank format) = null s - Strain (The name of the strain, will be used for annotated TALEs, OPTIONAL) = null outdir - The output directory, defaults to the current working directory (.) = .
You get a description of each tool by calling AnnoTALEcli-1.0.jar with the corresponding tool name and keyword "info", e.g.,
java -jar AnnoTALEcli-1.0.jar predict info
Output:
*TALE Prediction* predicts transcription activator-like effector (TALE) genes in an input sequence, typically a 'Xanthomonas' genome. 'TALE Prediction' is based in HMMer nucleotide HMM models that describe N-terminus, repeat region, and C-terminus of TALEs. The input 'Genome' may be provided in FastA or Genbank format. Optionally, you may provide a strain name that will be used in the temporary TALE names and names of output files. Regardless of the input format, 'TALE Prediction' generates output in Genbank format containing the annotations of TALE genes. If the original input has already been a Genbank file, TALE annotations are added to the existing ones. In addition, 'TALE Prediction' generates annotations in GFF format, and also outputs the DNA and AS sequences of the predicted TALEs in FastA format. 'TALE Prediction' tries hard to make the CDS annotation a proper gene model, starting from a start codon and ending with a Stop. If either start or stop codon are located within the originally predicted region that is homologous to TALE genes, this original hit region is still reported as mRNA. Putative pseudo genes, e.g., with premature stop codons, are marked accordingly. The TALE DNA sequences output of 'TALE Prediction' may serve as input of the 'TALE Analysis', 'TALE Class Builder', and 'TALE Class Assignment' tools. If you experience problems using 'TALE Prediction', please contact us.
Standard pipeline
Assuming that your current working directory contains the AnnoTALEcli Jar file, a genome of interest (of a hypothetical 'Xoo' strain PXO999 with accesion CP1234567) in a FastA file "genome.fa", all rice promoters in a FastA file "Rice-promoters.fa", and a directory "out" designated to hold all output files, a typical AnnoTALE pipeline could look like
java -jar AnnoTALEcli-1.0.jar predict g=genome.fa outdir=out java -jar AnnoTALEcli-1.0.jar analyze t=out/TALE_DNA_sequences.fasta outdir=out java -jar AnnoTALEcli-1.0.jar loadAndView outdir=out java -jar AnnoTALEcli-1.0.jar assign c=out/Class_builder_download.xml t=out/TALE_DNA_parts.fasta s="Xoo PXO999" a="CP1234567" outdir=out java -jar AnnoTALEcli-1.0.jar rename r=out/TALE_names_\(Xoo_PXO999\).tsv i=out/Genbank__TALE_predictions.gb outdir=out java -jar AnnoTALEcli-1.0.jar targets i=Rice-promoters.fa p="TALEs in class builder" c=out/Augmented_class_builder_\(Xoo_PXO999\).xml outdir=out
Afterwards, you find all output files of all those tools in the directory "out". The output files and directories are named in analogy to the names in the AnnoTALE GUI version (see User Guide for the GUI version)