 de.jstacs.models.hmm.AbstractHMM
de.jstacs.models.hmm.AbstractHMM
 de.jstacs.models.hmm.models.HigherOrderHMM
de.jstacs.models.hmm.models.HigherOrderHMM
|
||||||||||
| PREV CLASS NEXT CLASS | FRAMES NO FRAMES | |||||||||
| SUMMARY: NESTED | FIELD | CONSTR | METHOD | DETAIL: FIELD | CONSTR | METHOD | |||||||||
java.lang.Objectde.jstacs.models.hmm.AbstractHMM
de.jstacs.models.hmm.models.HigherOrderHMM
public class HigherOrderHMM
This class implements a higher order hidden Markov model.
Currently, the modeling of the transitions is higher order, but is easily possible to extend this to emissions.
This implementation allows to have a set of final states 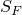 .
A state is denoted final states if it is allowed at the end of a path. Hence, any valid path always ends with a final state.
Using the method
.
A state is denoted final states if it is allowed at the end of a path. Hence, any valid path always ends with a final state.
Using the method AbstractHMM.getLogProbFor(Sequence) for sequence 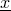 returns the value
returns the value
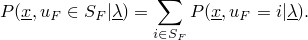
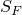 to all states leads to the computation of the likelihood.
to all states leads to the computation of the likelihood.
| Nested Class Summary | |
|---|---|
protected static class |
HigherOrderHMM.Type
This enum defined different types of computations that will be done using the backward algorithm. |
| Field Summary | |
|---|---|
protected double[] |
backwardIntermediate
Helper variable = only for internal use. |
protected int[] |
container
Helper variable = only for internal use. |
protected double[] |
logEmission
Helper variable = only for internal use. |
protected int[][] |
numberOfSummands
Helper variable = only for internal use. |
protected IntList |
stateList
Helper variable = only for internal use. |
| Fields inherited from class de.jstacs.models.hmm.AbstractHMM |
|---|
bwdMatrix, con, emission, emissionIdx, finalState, forward, fwdMatrix, name, sostream, START_NODE, states, trainingParameter, transition |
| Constructor Summary | |
|---|---|
HigherOrderHMM(HMMTrainingParameterSet trainingParameterSet,
String[] name,
Emission[] emission,
BasicHigherOrderTransition.AbstractTransitionElement... te)
This is a convenience constructor. |
|
HigherOrderHMM(HMMTrainingParameterSet trainingParameterSet,
String[] name,
int[] emissionIdx,
boolean[] forward,
Emission[] emission,
BasicHigherOrderTransition.AbstractTransitionElement... te)
This is the main constructor. |
|
HigherOrderHMM(StringBuffer xml)
The standard constructor for the interface Storable. |
|
| Method Summary | |
|---|---|
protected void |
appendFurtherInformation(StringBuffer xml)
This method appends further information to the XML representation. |
protected double |
baumWelch(int startPos,
int endPos,
double weight,
Sequence seq)
This method computes the likelihood and modifies the sufficient statistics according to the Baum-Welch algorithm. |
HigherOrderHMM |
clone()
Creates a clone (deep copy) of the current Model instance. |
protected void |
createHelperVariables()
This method instantiates all helper variables that are need inside the model for instance for filling forward and backward matrix, ... |
protected void |
createStates()
This method creates states for the internal usage. |
Sample |
emitSample(int numberOfSequences,
int... seqLength)
This method returns a Sample object containing artificial
sequence(s). |
protected void |
estimateFromStatistics()
This method estimates the parameters of all emissions and the transition using their sufficient statistics. |
protected void |
extractFurtherInformation(StringBuffer xml)
This method extracts further information from the XML representation. |
protected void |
fillBwdMatrix(int startPos,
int endPos,
Sequence seq)
This method fills the backward-matrix for a given sequence. |
protected void |
fillBwdOrViterbiMatrix(HigherOrderHMM.Type t,
int startPos,
int endPos,
double weight,
Sequence seq)
This method computes the entries of the backward or the viterbi matrix. |
protected void |
fillFwdMatrix(int startPos,
int endPos,
Sequence seq)
This method fills the forward-matrix for a given sequence. |
protected void |
fillLogStatePosteriorMatrix(double[][] statePosterior,
int startPos,
int endPos,
Sequence seq,
boolean silentZero)
This method fills the log state posterior of Sequence seq in a given matrix. |
protected void |
finalize()
|
ResultSet |
getCharacteristics()
Returns some information characterizing or describing the current instance of the model. |
String |
getInstanceName()
Should return a short instance name such as iMM(0), BN(2), ... |
int |
getLength()
Returns the length of sequences this model can classify. |
double |
getLogPriorTerm()
Returns a value that is proportional to the log of the prior. |
double[] |
getLogProbFor(Sample data)
This method computes the logarithm of the probabilities of all sequences in the given sample. |
void |
getLogProbFor(Sample data,
double[] res)
This method computes and stores the logarithm of the probabilities for any sequence in the sample in the given double-array. |
double |
getLogProbForPath(IntList path,
int startPos,
Sequence seq)
|
byte |
getMaximalMarkovOrder()
This method returns the maximal used Markov order, if possible. |
NumericalResultSet |
getNumericalCharacteristics()
Returns the subset of numerical values that are also returned by Model.getCharacteristics(). |
Pair<IntList,Double> |
getViterbiPathFor(int startPos,
int endPos,
Sequence seq)
|
protected String |
getXMLTag()
Returns the tag for the XML representation. |
protected void |
initializeRandomly()
This method initializes all emissions and the transition randomly. |
boolean |
isTrained()
Returns true if the model has been trained successfully,
false otherwise. |
protected void |
resetStatistics()
This method resets all sufficient statistics of all emissions and the transition. |
void |
samplePath(IntList path,
int startPos,
int endPos,
Sequence seq)
This method samples a valid path for the given sequence seq using the internal parameters. |
void |
train(Sample data,
double[] weights)
Trains the Model object given the data as Sample using
the specified weights. |
protected double |
viterbi(IntList path,
int startPos,
int endPos,
double weight,
Sequence seq)
This method computes the viterbi score of a given sequence seq. |
| Methods inherited from class java.lang.Object |
|---|
equals, getClass, hashCode, notify, notifyAll, wait, wait, wait |
| Field Detail |
|---|
protected int[] container
Transition.fillTransitionInformation(int, int, int, int[]).
protected double[] logEmission
AbstractHMM.emissionprotected double[] backwardIntermediate
numberOfSummands
protected int[][] numberOfSummands
protected IntList stateList
samplePath(IntList, int, int, Sequence).
| Constructor Detail |
|---|
public HigherOrderHMM(HMMTrainingParameterSet trainingParameterSet,
String[] name,
Emission[] emission,
BasicHigherOrderTransition.AbstractTransitionElement... te)
throws Exception
i used emission i on the forward strand.
trainingParameterSet - the ParameterSet that determines the training algorithm and contains the necessary Parametersname - the names of the statesemission - the emissionste - the BasicHigherOrderTransition.AbstractTransitionElements building a transition
Exception - if
name, emissionIdx, or forward is not equal to the number of statesAlphabetContainerHigherOrderHMM(HMMTrainingParameterSet, String[], int[], boolean[], Emission[], BasicHigherOrderTransition.AbstractTransitionElement...)
public HigherOrderHMM(HMMTrainingParameterSet trainingParameterSet,
String[] name,
int[] emissionIdx,
boolean[] forward,
Emission[] emission,
BasicHigherOrderTransition.AbstractTransitionElement... te)
throws Exception
trainingParameterSet - the ParameterSet that determines the training algorithm and contains the necessary Parametersname - the names of the statesemissionIdx - the indices of the emissions that should be used for each state, if null state i will use emission iforward - a boolean array that indicates whether the symbol on the forward or the reverse complementary strand should be used,
if null all states use the forward strandemission - the emissionste - the BasicHigherOrderTransition.AbstractTransitionElements building a transition
Exception - if
name, emissionIdx, or forward is not equal to the number of statesAlphabetContainer
public HigherOrderHMM(StringBuffer xml)
throws NonParsableException
Storable.
Constructs an HigherOrderHMM out of an XML representation.
xml - the XML representation as StringBuffer
NonParsableException - if the HigherOrderHMM could not be reconstructed out of
the StringBuffer xml| Method Detail |
|---|
protected void createHelperVariables()
AbstractHMM
createHelperVariables in class AbstractHMMprotected String getXMLTag()
AbstractHMM
getXMLTag in class AbstractHMMAbstractHMM.fromXML(StringBuffer),
AbstractHMM.toXML()protected void appendFurtherInformation(StringBuffer xml)
AbstractHMM
appendFurtherInformation in class AbstractHMMxml - the XML representation
protected void extractFurtherInformation(StringBuffer xml)
throws NonParsableException
extractFurtherInformation in class AbstractHMMxml - the XML representation
NonParsableException - if the information could not be reconstructed out of the StringBuffer xml
public HigherOrderHMM clone()
throws CloneNotSupportedException
ModelModel instance.
clone in interface Modelclone in class AbstractHMMCloneNotSupportedException - if something went wrong while cloningprotected void createStates()
AbstractHMM
createStates in class AbstractHMMpublic double getLogPriorTerm()
Model
Model.getPriorTerm()public int getLength()
Model
public double getLogProbForPath(IntList path,
int startPos,
Sequence seq)
throws Exception
getLogProbForPath in class AbstractHMMpath - the given state pathstartPos - the start position within the sequence(s) (inclusive)seq - the sequence(s)
Exception - if the probability for the sequence given path could not be computed, for instance if the model is not trained, ...
protected void fillLogStatePosteriorMatrix(double[][] statePosterior,
int startPos,
int endPos,
Sequence seq,
boolean silentZero)
throws Exception
AbstractHMMseq in a given matrix.
fillLogStatePosteriorMatrix in class AbstractHMMstatePosterior - the matrix for the log state posteriorstartPos - the start positionendPos - the end positionseq - the sequencesilentZero - true if the state posterior for silent states is defined to be zero, otherwise false
Exception - if an error occurs during the computationAbstractHMM.getLogStatePosteriorMatrixFor(int, int, Sequence),
AbstractHMM.createMatrixForStatePosterior(int, int)
protected void fillFwdMatrix(int startPos,
int endPos,
Sequence seq)
throws OperationNotSupportedException,
WrongLengthException
AbstractHMM
fillFwdMatrix in class AbstractHMMstartPos - the start position (inclusive) in the sequenceendPos - the end position (inclusive) in the sequenceseq - the sequence
OperationNotSupportedException
WrongLengthException
protected void fillBwdMatrix(int startPos,
int endPos,
Sequence seq)
throws Exception
AbstractHMM
fillBwdMatrix in class AbstractHMMstartPos - the start position (inclusive) in the sequenceendPos - the end position (inclusive) in the sequenceseq - the sequence
Exception - if some error occurs during the computation
protected void fillBwdOrViterbiMatrix(HigherOrderHMM.Type t,
int startPos,
int endPos,
double weight,
Sequence seq)
throws Exception
t - a switch to decide which computation modestartPos - start position of the sequenceendPos - end position of the sequenceweight - the given external weight of the sequence (only used for Baum-Welch)seq - the sequence
Exception - forwarded from TrainableState.addToStatistic(int, int, double, de.jstacs.data.Sequence) and State.getLogScoreFor(int, int, Sequence)
public Pair<IntList,Double> getViterbiPathFor(int startPos,
int endPos,
Sequence seq)
throws Exception
getViterbiPathFor in class AbstractHMMstartPos - the start position within the sequenceendPos - the end position within the sequenceseq - the sequence
Pair containing the viterbi state path and the corresponding score
Exception - if the viterbi path could not be computed, for instance if the model is not trained, ...
protected double viterbi(IntList path,
int startPos,
int endPos,
double weight,
Sequence seq)
throws Exception
seq.
Furthermore, it allows either to modify the sufficient statistics according
to the viterbi training algorithm or to compute the viterbi path, which will
in this case be returned in path.
path - if null viterbi training, otherwise computation of the viterbi pathstartPos - the start positionendPos - the end positionweight - the sequence weight, in most cases this is 1seq - the sequence
Exception - an error occurs during the computation
protected double baumWelch(int startPos,
int endPos,
double weight,
Sequence seq)
throws Exception
startPos - the start positionendPos - the end positionweight - the sequence weight, in most cases this is 1seq - the sequence
Exception - an error occurs during the computation
public void train(Sample data,
double[] weights)
throws Exception
ModelModel object given the data as Sample using
the specified weights. The weight at position i belongs to the element at
position i. So the array weight should have the number of
sequences in the sample as dimension. (Optionally it is possible to use
weight == null if all weights have the value one.)train(data1); train(data2)
should be a fully trained model over data2 and not over
data1+data2. All parameters of the model were given by the
call of the constructor.
data - the given sequences as Sampleweights - the weights of the elements, each weight should be
non-negative
Exception - if the training did not succeed (e.g. the dimension of
weights and the number of sequences in the
sample do not match)Sample.getElementAt(int),
Sample.ElementEnumeratorprotected void initializeRandomly()
protected void resetStatistics()
protected void estimateFromStatistics()
public final byte getMaximalMarkovOrder()
throws UnsupportedOperationException
Model
UnsupportedOperationException - if the model can't give a proper answer
public ResultSet getCharacteristics()
throws Exception
ModelStorableResult.
Exception - if some of the characteristics could not be definedStorableResultpublic String getInstanceName()
Model
public double[] getLogProbFor(Sample data)
throws Exception
ModelModel.getLogProbFor(Sequence).
data - the sample of sequences
Exception - if something went wrongModel.getLogProbFor(Sequence)
public void getLogProbFor(Sample data,
double[] res)
throws Exception
Modeldouble-array.
Model.getLogProbFor(Sequence).
data - the sample of sequencesres - the array for the results, has to have length
data.getNumberOfElements() (which returns the
number of sequences in the sample)
Exception - if something went wrongModel.getLogProbFor(Sample)
public Sample emitSample(int numberOfSequences,
int... seqLength)
throws NotTrainedException,
Exception
ModelSample object containing artificial
sequence(s).
emitSample( int n, int l ) should return a sample with
n sequences of length l.
emitSample( int n, int[] l ) should return a sample with
n sequences which have a sequence length corresponding to
the entry in the given array l.
emitSample( int n ) and
emitSample( int n, null ) should return a sample with
n sequences of length of the model (
Model.getLength()).
Exception.
numberOfSequences - the number of sequences that should be contained in the
returned sampleseqLength - the length of the sequences for a homogeneous model; for an
inhomogeneous model this parameter should be null
or an array of size 0.
Sample containing the artificial sequence(s)
NotTrainedException - if the model is not trained yet
Exception - if the emission did not succeedSample
public NumericalResultSet getNumericalCharacteristics()
throws Exception
ModelModel.getCharacteristics().
Exception - if some of the characteristics could not be definedpublic boolean isTrained()
Modeltrue if the model has been trained successfully,
false otherwise.
true if the model has been trained successfully,
false otherwise
protected void finalize()
throws Throwable
finalize in class AbstractHMMThrowable
public void samplePath(IntList path,
int startPos,
int endPos,
Sequence seq)
throws Exception
seq using the internal parameters.
path - an IntList containing the path after using this methodstartPos - the start positionendPos - the end positionseq - the sequence
Exception - if an error occurs during computation
|
||||||||||
| PREV CLASS NEXT CLASS | FRAMES NO FRAMES | |||||||||
| SUMMARY: NESTED | FIELD | CONSTR | METHOD | DETAIL: FIELD | CONSTR | METHOD | |||||||||