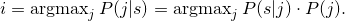- java.lang.Object
-
- de.jstacs.sequenceScores.statisticalModels.trainable.AbstractTrainableStatisticalModel
-
- de.jstacs.sequenceScores.statisticalModels.trainable.mixture.AbstractMixtureTrainSM
-
- All Implemented Interfaces:
- SequenceScore, StatisticalModel, TrainableStatisticalModel, Storable, Cloneable
- Direct Known Subclasses:
- HiddenMotifMixture, MixtureTrainSM, StrandTrainSM
public abstract class AbstractMixtureTrainSM extends AbstractTrainableStatisticalModel
This is the abstract class for all kinds of mixture models. It enables the user to train the parameters usingAbstractMixtureTrainSM.Algorithm.EMorAbstractMixtureTrainSM.Algorithm.GIBBS_SAMPLING. If this instance is trained usingAbstractMixtureTrainSM.Algorithm.GIBBS_SAMPLINGthe internal models that will be adjusted have to implementSamplingComponent. If you use Gibbs sampling temporary files will be created in the Java temp folder. These files will be deleted if no reference to the current instance exists and the Garbage Collector is called. Therefore it is recommended to call the Garbage Collector explicitly at the end of any application.
The model stores a reference to the last data set used intrain. This enables the user to estimate the parameters iteratively beginning with the current set of parameters. To this end, you can use the methodcontinueIterations(double[], double[][], int, int).
The methodsetOutputStream(OutputStream)enables the user to get comments from thetrain(DataSet, double[])method or to repress them.
The methodgetScoreForBestRun()enables the user to optimize different instances of the same model (clone()) using the EM-algorithm on different CPUs, to compare the results and to select the best trained model. This might be useful to get the results faster (measured in real time).
The reference to the internal data set is not stored if the model is stored in aStringBuffer. So you can use these methods only after training the parameters after (re)creating a model.- Author:
- Jens Keilwagen, Berit Haldemann
- See Also:
SamplingComponent,System.gc()
-
-
Nested Class Summary
Nested Classes Modifier and Type Class and Description static classAbstractMixtureTrainSM.AlgorithmThisenumdefines the different types of algorithms that can be used in anAbstractMixtureTrainSM.static classAbstractMixtureTrainSM.ParameterizationThisenumdefines the different types of parameterization for a probability that can be used in anAbstractMixtureTrainSM.
-
Field Summary
Fields Modifier and Type Field and Description protected AbstractMixtureTrainSM.AlgorithmalgorithmThe type of algorithm.protected booleanalgorithmHasBeenRunA switch which indicates that the algorithm for determining the parameters has been run.protected TrainableStatisticalModel[]alternativeModelThe alternative models for the EM.protected doublebestThis field contains the value of objective function of the best start of the training.protected BurnInTestburnInTestTheBurnInTestthat is used to stop the sampling.protected double[]componentHyperParamsThe hyperparameters for estimating the probabilities of the components.protected double[]compProbThis array is used while training to avoid creating many new objects.protected int[]counterThe current index of the parameter set while adjustment (optimization).protected intdimensionThe number of dimensions.protected booleanestimateComponentProbsThe switch for estimating the component probabilities or not.protected File[]fileThe file in which the component probabilities are stored.protected BufferedReaderfilereaderReading component probabilities from a file.protected BufferedWriterfilewriterSaving component probabilities in a file.protected intinitialIterationThe number of initial iterations.protected double[]logWeightsThe log probabilities for each component.protected TrainableStatisticalModel[]modelThe model for the sequences.protected boolean[]optimizeModelA switch for each model whether to optimize/adjust or not.protected DataSet[]sampleThe data set that was used in the last training.protected intsamplingIndexThe current index of the sampling.protected double[][]seqWeightsThe weights of the (sub-)sequence used to train the components (internal models).protected SafeOutputStreamsostreamThis is the stream for writing information while training.protected intstartsThe number of starts.protected intstationaryIterationThe number of (stationary) iterations of the Gibbs Sampler.protected double[]weightsThe probabilities for each component.-
Fields inherited from class de.jstacs.sequenceScores.statisticalModels.trainable.AbstractTrainableStatisticalModel
alphabets, length
-
-
Constructor Summary
Constructors Modifier Constructor and Description protectedAbstractMixtureTrainSM(int length, TrainableStatisticalModel[] models, boolean[] optimizeModel, int dimension, int starts, boolean estimateComponentProbs, double[] componentHyperParams, double[] weights, AbstractMixtureTrainSM.Algorithm algorithm, double alpha, TerminationCondition tc, AbstractMixtureTrainSM.Parameterization parametrization, int initialIteration, int stationaryIteration, BurnInTest burnInTest)Creates a newAbstractMixtureTrainSM.protectedAbstractMixtureTrainSM(StringBuffer xml)The standard constructor for the interfaceStorable.
-
Method Summary
All Methods Static Methods Instance Methods Abstract Methods Concrete Methods Modifier and Type Method and Description booleanalgorithmHasBeenRun()This method indicates whether the parameters of the model has been determined by the internal algorithm.protected voidcheckLength(int index, int l)This method checks if the lengthlof the model with indexindexis capable for the current instance.protected voidcheckModelsForGibbsSampling()This method can be used to check whether the necessary models have implemented theSamplingComponent.AbstractMixtureTrainSMclone()Follows the conventions ofObject'sclone()-method.protected doublecontinueIterations(double[] dataWeights, double[][] seqweights)This method will run the train algorithm for the current model on the internal data set.protected doublecontinueIterations(double[] dataWeights, double[][] seqweights, int iterations, int start)This method will run the train algorithm for the current model on the internal sample.protected double[][]createSeqWeightsArray()Creates an array that can be used for weighting sequences in the algorithm.protected double[][]doFirstIteration(DataSet data, double[] dataWeights)This method will do the first step in the train algorithm for the current model.protected double[][]doFirstIteration(DataSet data, double[] dataWeights, MultivariateRandomGenerator m, MRGParams[] params)This method will do the first step in the train algorithm for the current model.protected abstract double[][]doFirstIteration(double[] dataWeights, MultivariateRandomGenerator m, MRGParams[] params)This method will do the first step in the train algorithm for the current model on the internal data set.static intdraw(double[] w, int start)This method draws an index of an array corresponding to the probabilities encoded in the entries of the array.DataSetemitDataSet(int n, int... lengths)This method returns aDataSetobject containing artificial sequence(s).protected abstract Sequence[]emitDataSetUsingCurrentParameterSet(int n, int... lengths)The method returns an array of sequences using the current parameter set.protected voidextendSampling(int sampling)This method prepares the model to extend an existing sampling.protected voidextractFurtherInformation(StringBuffer xml)This method is used in the subclasses to extract further information from the XML representation and to set these as values of the instance.protected voidfinalize()protected voidfromXML(StringBuffer representation)This method should only be used by the constructor that works on aStringBuffer.ResultSetgetCharacteristics()Returns some information characterizing or describing the current instance.protected StringBuffergetFurtherInformation()This method is used in the subclasses to append further information to the XML representation.intgetIndexOfMaximalComponentFor(Sequence s)Returns the indexiof the component withP(i|s)maximal.StringgetInstanceName()Should return a short instance name such as iMM(0), BN(2), ...doublegetLogPriorTerm()Returns a value that is proportional to the log of the prior.protected doublegetLogPriorTermForComponentProbs()This method computes the part of the prior that comes from the component probabilities.doublegetLogProbFor(int component, Sequence s)Returns the logarithmic probability for the sequence and the given component.doublegetLogProbFor(int component, Sequence s, int start, int end)Returns the logarithmic probability for the sequence between start and end and the given component.doublegetLogProbFor(Sequence sequence, int startpos, int endpos)Returns the logarithm of the probability of (a part of) the given sequence given the model.protected abstract doublegetLogProbUsingCurrentParameterSetFor(int component, Sequence s, int start, int end)Returns the logarithmic probability for the sequence and the given component using the current parameter set.double[]getLogScoreFor(DataSet data)This method computes the logarithm of the scores of all sequences in the given data set.TrainableStatisticalModelgetModel(int i)Returns a deep copy of thei-th model.TrainableStatisticalModel[]getModels()Returns a deep copy of the models.protected MultivariateRandomGeneratorgetMRG()This method creates the multivariate random generator that will be used during initialization.protected MRGParamsgetMRGParams()This method creates the parameters used in a multivariate random generator while initialization.StringgetNameOfAlgorithm()Returns the name of the used algorithm.protected voidgetNewComponentProbs(double[] weights)Estimates the weights of each component.protected voidgetNewParameters(int iteration, double[][] seqWeights, double[] w)This method trains the internal models on the internal data set and the given weights.protected voidgetNewParametersForModel(int modelIndex, int iteration, int sampleIndex, double[] seqWeights)This method trains the internal model with indexmodelIndexon the internal data set and the given weights.protected abstract doublegetNewWeights(double[] dataWeights, double[] w, double[][] seqweights)Computes sequence weights and returns the score.intgetNumberOfComponents()Returns the number of components the are modeled by thisAbstractMixtureTrainSM.NumericalResultSetgetNumericalCharacteristics()Returns the subset of numerical values that are also returned bySequenceScore.getCharacteristics().doublegetScoreForBestRun()Returns the value of the optimized function from the best run of the last training.double[]getWeights()This method returns a deep copy of the weights for each component.protected voidinitModelForSampling(int starts)This method initializes the model for the sampling.protected voidinitWithPrior(double[] w)This method sets the initial weights before counting the usage of each component.booleanisInitialized()This method can be used to determine whether the instance is initialized.protected booleanisInSamplingMode()This method returnstrueif the object is currently used in a sampling, otherwisefalse.doubleiterate(DataSet data, double[] dataWeights, MultivariateRandomGenerator m, MRGParams[] params)This method runs the train algorithm for the current model.protected doubleiterate(int start, double[] dataWeights, MultivariateRandomGenerator m, MRGParams[] params)This method runs the train algorithm for the current model and the internal data set.static intmax(double[] w, int start, int end)This method returns the index of a maximal entry in the arraywbetween indexstartandend.protected doublemodifyWeights(double[] w)This method modifies the computed weights for one sequence and returns the score.protected booleanparseNextParameterSet()This method allows the user to parse the next set of parameters (from a file).protected booleanparseParameterSet(int sampling, int burnInIteration)This method allows the user to parse the set of parameters with indexburnInIterationof a specificsampling(from a file).protected voidsamplingStopped()This method is the opposite of the methodinitModelForSampling(int).voidsetAlpha(double alpha)Sets the parameter of the Dirichlet distribution which is used when you invoketrainto init the gammas.voidsetOutputStream(OutputStream o)Sets theOutputStreamthat is used e.g.protected abstract voidsetTrainData(DataSet data)This method is invoked by thetrain-method and sets for a given data set the data set that should be used fortrain.protected voidsetWeights(double... weights)Sets the weights of each component.protected voidswap()This method swaps the current component models with the alternative model.StringBuffertoXML()This method returns an XML representation asStringBufferof an instance of the implementing class.voidtrain(DataSet data, double[] dataWeights)Trains theTrainableStatisticalModelobject given the data asDataSetusing the specified weights.-
Methods inherited from class de.jstacs.sequenceScores.statisticalModels.trainable.AbstractTrainableStatisticalModel
check, getAlphabetContainer, getLength, getLogProbFor, getLogProbFor, getLogScoreFor, getLogScoreFor, getLogScoreFor, getLogScoreFor, getMaximalMarkovOrder, toString, train
-
Methods inherited from class java.lang.Object
equals, getClass, hashCode, notify, notifyAll, wait, wait, wait
-
Methods inherited from interface de.jstacs.sequenceScores.SequenceScore
toString
-
-
-
-
Field Detail
-
weights
protected double[] weights
The probabilities for each component.
-
logWeights
protected double[] logWeights
The log probabilities for each component.
-
componentHyperParams
protected double[] componentHyperParams
The hyperparameters for estimating the probabilities of the components.
-
model
protected TrainableStatisticalModel[] model
The model for the sequences.
-
alternativeModel
protected TrainableStatisticalModel[] alternativeModel
The alternative models for the EM.
-
starts
protected int starts
The number of starts.
-
dimension
protected int dimension
The number of dimensions.
-
best
protected double best
This field contains the value of objective function of the best start of the training.
-
sostream
protected SafeOutputStream sostream
This is the stream for writing information while training.
-
sample
protected DataSet[] sample
The data set that was used in the last training. Will not be stored in theStringBufferwhen invokingtoXML().
-
estimateComponentProbs
protected boolean estimateComponentProbs
The switch for estimating the component probabilities or not.
-
optimizeModel
protected boolean[] optimizeModel
A switch for each model whether to optimize/adjust or not.
-
algorithm
protected AbstractMixtureTrainSM.Algorithm algorithm
The type of algorithm.
-
algorithmHasBeenRun
protected boolean algorithmHasBeenRun
A switch which indicates that the algorithm for determining the parameters has been run.
-
initialIteration
protected int initialIteration
The number of initial iterations.
-
stationaryIteration
protected int stationaryIteration
The number of (stationary) iterations of the Gibbs Sampler.
-
burnInTest
protected BurnInTest burnInTest
TheBurnInTestthat is used to stop the sampling.
-
filewriter
protected BufferedWriter filewriter
Saving component probabilities in a file.
-
filereader
protected BufferedReader filereader
Reading component probabilities from a file.
-
file
protected File[] file
The file in which the component probabilities are stored.
-
counter
protected int[] counter
The current index of the parameter set while adjustment (optimization).
-
samplingIndex
protected int samplingIndex
The current index of the sampling.
-
compProb
protected double[] compProb
This array is used while training to avoid creating many new objects.
-
seqWeights
protected double[][] seqWeights
The weights of the (sub-)sequence used to train the components (internal models). The first dimension is used for the models, the second for the (sub-)sequences.
-
-
Constructor Detail
-
AbstractMixtureTrainSM
protected AbstractMixtureTrainSM(int length, TrainableStatisticalModel[] models, boolean[] optimizeModel, int dimension, int starts, boolean estimateComponentProbs, double[] componentHyperParams, double[] weights, AbstractMixtureTrainSM.Algorithm algorithm, double alpha, TerminationCondition tc, AbstractMixtureTrainSM.Parameterization parametrization, int initialIteration, int stationaryIteration, BurnInTest burnInTest) throws CloneNotSupportedException, IllegalArgumentException, WrongAlphabetExceptionCreates a newAbstractMixtureTrainSM. This constructor can be used for any algorithm since it takes all necessary values as parameters.- Parameters:
length- the length used in this modelmodels- the single models building theAbstractMixtureTrainSM, if the model is trained usingAbstractMixtureTrainSM.Algorithm.GIBBS_SAMPLINGthe models that will be adjusted have to implementSamplingComponentoptimizeModel- an array of switches to determine whether a model should be optimized or notdimension- the number of componentsstarts- the number of times the algorithm will be started in thetrain-method, at least 1estimateComponentProbs- the switch for estimating the component probabilities in the algorithm or to hold them fixed; if the component parameters are fixed, the values ofweightswill be used, otherwise thecomponentHyperParamswill be incorporated in the adjustmentcomponentHyperParams- the hyperparameters for the component assignment prior- will only be used if
estimateComponentProbs == true - the array has to be
nullor has to have lengthdimension nullor an array with all values zero (0) then ML- otherwise (all values positive) a prior is used (MAP, MP, ...)
- depends on the
parameterization
- will only be used if
weights-nullor the weights for the components (thenweights.length == dimension)algorithm- eitherAbstractMixtureTrainSM.Algorithm.EMorAbstractMixtureTrainSM.Algorithm.GIBBS_SAMPLINGalpha- only forAbstractMixtureTrainSM.Algorithm.EM
the positive parameter for the Dirichlet distribution which is used when you invoketrainto initialize the gammas. It is recommended to usealpha = 1(uniform distribution on a simplex).tc- only forAbstractMixtureTrainSM.Algorithm.EM
theTerminationConditionfor stopping the EM-algorithm,tchas to returntruefromTerminationCondition.isSimple()parametrization- only forAbstractMixtureTrainSM.Algorithm.EM
the type of the component probability parameterization;AbstractMixtureTrainSM.Parameterization.THETAorAbstractMixtureTrainSM.Parameterization.LAMBDA- the parameterization of a component is determined by the component model
- it is recommended to use the same parameterization for the components and the component assignment probabilities
- it is recommended to use
AbstractMixtureTrainSM.Parameterization.LAMBDA
initialIteration- only forAbstractMixtureTrainSM.Algorithm.GIBBS_SAMPLINGthe positive length of the initial sampling phase (at least 1, at moststationaryIteration/starts)stationaryIteration- only forAbstractMixtureTrainSM.Algorithm.GIBBS_SAMPLINGthe positive length of the stationary phase (at least 1) (summed over all starts), i.e. the number of parameter sets that is used for approximationburnInTest- only forAbstractMixtureTrainSM.Algorithm.GIBBS_SAMPLINGthe test that will be used to determine the length of the burn-in phase- Throws:
IllegalArgumentException- if- the models are not able to score the sequence of length
length dimension < 1-
weights != null && weights.length != dimension weights != nulland it exists aniwhereweights[i] < 0starts < 1componentHyperParamsare not correct- the algorithm specific parameters are not correct
- the models are not able to score the sequence of length
WrongAlphabetException- if not allmodelswork on the same alphabetCloneNotSupportedException- if themodelscan not be cloned
-
AbstractMixtureTrainSM
protected AbstractMixtureTrainSM(StringBuffer xml) throws NonParsableException
The standard constructor for the interfaceStorable. Creates a newAbstractMixtureTrainSMout of its XML representation.- Parameters:
xml- the XML representation of the model asStringBuffer- Throws:
NonParsableException- if theStringBuffercan not be parsed
-
-
Method Detail
-
clone
public AbstractMixtureTrainSM clone() throws CloneNotSupportedException
Description copied from class:AbstractTrainableStatisticalModelFollows the conventions ofObject'sclone()-method.- Specified by:
clonein interfaceSequenceScore- Specified by:
clonein interfaceTrainableStatisticalModel- Overrides:
clonein classAbstractTrainableStatisticalModel- Returns:
- an object, that is a copy of the current
AbstractTrainableStatisticalModel(the member-AlphabetContainerisn't deeply cloned since it is assumed to be immutable). The type of the returned object is defined by the classXdirectly inherited fromAbstractTrainableStatisticalModel. HenceX'sclone()-method should work as:
1.Object o = (X)super.clone();
2. all additional member variables ofodefined byXthat are not of simple data-types likeint,double, ... have to be deeply copied
3.return o - Throws:
CloneNotSupportedException- if something went wrong while cloning
-
getMRG
protected MultivariateRandomGenerator getMRG()
This method creates the multivariate random generator that will be used during initialization.- Returns:
- a multivariate random generator
- See Also:
getMRGParams()
-
getMRGParams
protected MRGParams getMRGParams()
This method creates the parameters used in a multivariate random generator while initialization.- Returns:
- the parameters for the multivariate random generator
- See Also:
getMRG()
-
train
public void train(DataSet data, double[] dataWeights) throws Exception
Description copied from interface:TrainableStatisticalModelTrains theTrainableStatisticalModelobject given the data asDataSetusing the specified weights. The weight at position i belongs to the element at position i. So the arrayweightshould have the number of sequences in the data set as dimension. (Optionally it is possible to useweight == nullif all weights have the value one.)
This method should work non-incrementally. That means the result of the following series:train(data1);train(data2)should be a fully trained model overdata2and not overdata1+data2. All parameters of the model were given by the call of the constructor.- Parameters:
data- the given sequences asDataSetdataWeights- the weights of the elements, each weight should be non-negative- Throws:
Exception- if the training did not succeed (e.g. the dimension ofweightsand the number of sequences in the data set do not match)- See Also:
DataSet.getElementAt(int),DataSet.ElementEnumerator
-
swap
protected void swap()
This method swaps the current component models with the alternative model.
This method should NOT be made public and should ONLY be used in thetrain-method.
-
setTrainData
protected abstract void setTrainData(DataSet data) throws Exception
This method is invoked by thetrain-method and sets for a given data set the data set that should be used fortrain.- Parameters:
data- the given data set of sequences- Throws:
Exception- if something went wrong
-
createSeqWeightsArray
protected double[][] createSeqWeightsArray()
Creates an array that can be used for weighting sequences in the algorithm.- Returns:
- an array that can be used for weighting sequences in the algorithm
-
iterate
public double iterate(DataSet data, double[] dataWeights, MultivariateRandomGenerator m, MRGParams[] params) throws Exception
This method runs the train algorithm for the current model.- Parameters:
data- the data set of sequencesdataWeights- the weights for each sequence ornullm- the random generator for initiating the algorithmparams- the parameters for the sequences- Returns:
- the score
- Throws:
Exception- if something went wrong- See Also:
doFirstIteration(DataSet, double[], MultivariateRandomGenerator, MRGParams[]),continueIterations(double[], double[][]),continueIterations(double[], double[][], int, int)
-
iterate
protected double iterate(int start, double[] dataWeights, MultivariateRandomGenerator m, MRGParams[] params) throws ExceptionThis method runs the train algorithm for the current model and the internal data set.- Parameters:
start- the index of the trainingdataWeights- the weights for each sequence ornullm- the random generator for initiating the algorithmparams- the parameters for the sequences- Returns:
- the score
- Throws:
Exception- if something went wrong- See Also:
doFirstIteration(DataSet, double[], MultivariateRandomGenerator, MRGParams[]),continueIterations(double[], double[][]),continueIterations(double[], double[][], int, int)
-
doFirstIteration
protected double[][] doFirstIteration(DataSet data, double[] dataWeights) throws Exception
This method will do the first step in the train algorithm for the current model. The initialization will be done by randomly setting the component membership. This is useful when nothing is known about the problem.- Parameters:
data- the data set of sequencesdataWeights-nullor the weights of each element of the data set- Returns:
- the weighting array used to initialize, this array can be reused in the following iterations
- Throws:
Exception- if something went wrong
-
doFirstIteration
protected double[][] doFirstIteration(DataSet data, double[] dataWeights, MultivariateRandomGenerator m, MRGParams[] params) throws Exception
This method will do the first step in the train algorithm for the current model. The initialization will be done by randomly setting the component membership. This is useful when nothing is known about the problem.- Parameters:
data- the data set of sequencesdataWeights-nullor the weights of each element of the data setm- the multivariate random generatorparams- the parameters for the multivariate random generator- Returns:
- the weighting array used to initialize, this array can be reused in the following iterations
- Throws:
Exception- if something went wrong
-
doFirstIteration
protected abstract double[][] doFirstIteration(double[] dataWeights, MultivariateRandomGenerator m, MRGParams[] params) throws ExceptionThis method will do the first step in the train algorithm for the current model on the internal data set. The initialization will be done by randomly setting the component membership. This is useful when nothing is known about the problem.- Parameters:
dataWeights-nullor the weights of each element of the data setm- the multivariate random generatorparams- the parameters for the multivariate random generator- Returns:
- the weighting array used to initialize, this array can be reused in the following iterations
- Throws:
Exception- if something went wrong
-
continueIterations
protected double continueIterations(double[] dataWeights, double[][] seqweights) throws ExceptionThis method will run the train algorithm for the current model on the internal data set. The initialization will be done by using the models of theAbstractMixtureTrainSM. So in this case the models have to be trained already. This method is useful for restarting the train algorithm at a certain point. The algorithm will stop if the difference between the optimized functions for two iterations is smaller than the specified threshold.
If the difference becomes significant negative an exception is thrown.- Parameters:
dataWeights-nullor the weights of each element of the internal data set (last data set theAbstractMixtureTrainSMwas trained on)seqweights-nullor an array for weighting the sequences, seecreateSeqWeightsArray()- Returns:
- a score for the model
- Throws:
Exception- if something went wrong
-
continueIterations
protected double continueIterations(double[] dataWeights, double[][] seqweights, int iterations, int start) throws ExceptionThis method will run the train algorithm for the current model on the internal sample. The initialization will be done by using the models of theAbstractMixtureTrainSM. So in this case the models have to be trained already. This method is useful for restarting the algorithm at a certain point. The algorithm will stop after the number of iterations.- Parameters:
dataWeights-nullor the weights of each element of the internal sample (last sample theAbstractMixtureTrainSMwas trained on)seqweights-nullor an array for weighting the sequences, seecreateSeqWeightsArray()iterations- the number of iterations that should be donestart- the index of the run in aTrainableStatisticalModel.train(DataSet)-call- Returns:
- the current score (likelihood or posterior)
- Throws:
Exception- if something went wrong
-
getNewParameters
protected void getNewParameters(int iteration, double[][] seqWeights, double[] w) throws ExceptionThis method trains the internal models on the internal data set and the given weights.- Parameters:
iteration- the number of times this method has been invokedseqWeights- the weights for each model and sequencew- the weights for the components- Throws:
Exception- if the training of the internal models went wrong
-
getNewParametersForModel
protected void getNewParametersForModel(int modelIndex, int iteration, int sampleIndex, double[] seqWeights) throws ExceptionThis method trains the internal model with indexmodelIndexon the internal data set and the given weights.- Parameters:
modelIndex- the index of the modeliteration- the number of times this method has been invoked for this modelsampleIndex- the index of the internal data set that should be usedseqWeights- the weights for each sequence- Throws:
Exception- if the training of the internal model went wrong
-
getNewWeights
protected abstract double getNewWeights(double[] dataWeights, double[] w, double[][] seqweights) throws ExceptionComputes sequence weights and returns the score.- Parameters:
dataWeights- the weights for the internal data set (should not be changed)w- the array for the statistic of the component parameters (shall be filled)seqweights- an array containing for each component the weights for each sequence (shall be filled)- Returns:
- the score
- Throws:
Exception- if something went wrong
-
modifyWeights
protected double modifyWeights(double[] w)
This method modifies the computed weights for one sequence and returns the score.- Parameters:
w- the weights- Returns:
- the score
-
initWithPrior
protected void initWithPrior(double[] w)
This method sets the initial weights before counting the usage of each component. For ML the weights are set to 0 and for MAP they are set to the component hyperparameters.- Parameters:
w- the array of weights
-
getLogProbFor
public double getLogProbFor(int component, Sequence s) throws ExceptionReturns the logarithmic probability for the sequence and the given component.- Parameters:
component- the index of the components- the sequence- Returns:
log P(s,component) = log P(s|component) + log P(component)- Throws:
Exception- if the model was not trained yet or something else went wrong- See Also:
getNumberOfComponents()
-
getLogProbFor
public double getLogProbFor(int component, Sequence s, int start, int end) throws ExceptionReturns the logarithmic probability for the sequence between start and end and the given component.- Parameters:
component- the index of the components- the sequencestart- the start position in the sequenceend- the end position in the sequence- Returns:
log P(s[start..end],component) = log P(s[start..end]|component) + log P(component)- Throws:
Exception- if the model was not trained yet or something else went wrong- See Also:
getNumberOfComponents()
-
getLogProbUsingCurrentParameterSetFor
protected abstract double getLogProbUsingCurrentParameterSetFor(int component, Sequence s, int start, int end) throws ExceptionReturns the logarithmic probability for the sequence and the given component using the current parameter set.- Parameters:
component- the index of the components- the sequencestart- the start position in the sequenceend- the end position in the sequence- Returns:
log P(s,component) = log P(s|component) + log P(component)- Throws:
Exception- if not trained yet or something else went wrong- See Also:
getNumberOfComponents()
-
getLogProbFor
public final double getLogProbFor(Sequence sequence, int startpos, int endpos) throws Exception
Description copied from interface:StatisticalModelReturns the logarithm of the probability of (a part of) the given sequence given the model. If at least one random variable is continuous the value of density function is returned.
It extends the possibility given by the methodStatisticalModel.getLogProbFor(Sequence, int)by the fact, that the model could be e.g. homogeneous and therefore the length of the sequences, whose probability should be returned, is not fixed. Additionally, the end position of the part of the given sequence is given and the probability of the part from positionstartpostoendpos(inclusive) should be returned.
Thelengthand thealphabetsdefine the type of data that can be modeled and therefore both has to be checked.- Parameters:
sequence- the given sequencestartpos- the start position within the given sequenceendpos- the last position to be taken into account- Returns:
- the logarithm of the probability or the value of the density function of (the part of) the given sequence given the model
- Throws:
Exception- if the sequence could not be handled (e.g.startpos >,endpos > sequence.length, ...) by the modelNotTrainedException- if the model is not trained yet
-
getLogScoreFor
public final double[] getLogScoreFor(DataSet data) throws Exception
Description copied from interface:SequenceScoreThis method computes the logarithm of the scores of all sequences in the given data set. The values are stored in an array according to the index of the respective sequence in the data set.
The score for any sequence shall be computed independent of all other sequences in the data set. So the result should be exactly the same as for the methodSequenceScore.getLogScoreFor(Sequence).- Specified by:
getLogScoreForin interfaceSequenceScore- Overrides:
getLogScoreForin classAbstractTrainableStatisticalModel- Parameters:
data- the data set of sequences- Returns:
- an array containing the logarithm of the score of all sequences of the data set
- Throws:
Exception- if something went wrong- See Also:
SequenceScore.getLogScoreFor(Sequence)
-
getLogPriorTerm
public double getLogPriorTerm() throws ExceptionDescription copied from interface:StatisticalModelReturns a value that is proportional to the log of the prior. For maximum likelihood (ML) 0 should be returned.- Returns:
- a value that is proportional to the log of the prior
- Throws:
Exception- if something went wrong
-
getLogPriorTermForComponentProbs
protected final double getLogPriorTermForComponentProbs()
This method computes the part of the prior that comes from the component probabilities.- Returns:
- the part of the prior that comes from the component probabilities
-
getScoreForBestRun
public final double getScoreForBestRun() throws NotTrainedException, OperationNotSupportedExceptionReturns the value of the optimized function from the best run of the last training.- Returns:
- the value of the optimized function from the best run of the last training
- Throws:
NotTrainedException- if the training algorithm has not been runOperationNotSupportedException- if this method is used for an instance that does not use the EM- See Also:
train(DataSet, double[]),algorithmHasBeenRun()
-
getInstanceName
public String getInstanceName()
Description copied from interface:SequenceScoreShould return a short instance name such as iMM(0), BN(2), ...- Returns:
- a short instance name
-
getIndexOfMaximalComponentFor
public int getIndexOfMaximalComponentFor(Sequence s) throws Exception
Returns the indexiof the component withP(i|s)maximal. Therefore it computes- Parameters:
s- the sequence- Returns:
- the index of the component
- Throws:
Exception- if the model was not trained yet or something else went wrong- See Also:
getLogProbFor(int, Sequence)
-
getModels
public final TrainableStatisticalModel[] getModels() throws CloneNotSupportedException
Returns a deep copy of the models.- Returns:
- an array of
AbstractTrainableStatisticalModels - Throws:
CloneNotSupportedException- if at least one model can not be cloned- See Also:
getModel(int)
-
getModel
public final TrainableStatisticalModel getModel(int i) throws CloneNotSupportedException
Returns a deep copy of thei-th model.- Parameters:
i- the index- Returns:
- a deep copy of the
i-th model - Throws:
CloneNotSupportedException- if at least one model can not be cloned- See Also:
getModels()
-
getNameOfAlgorithm
public String getNameOfAlgorithm()
Returns the name of the used algorithm.- Returns:
- the name of the used algorithm
-
getNumberOfComponents
public final int getNumberOfComponents()
Returns the number of components the are modeled by thisAbstractMixtureTrainSM.- Returns:
- the number of components
-
getCharacteristics
public ResultSet getCharacteristics() throws Exception
Description copied from interface:SequenceScoreReturns some information characterizing or describing the current instance. This could be e.g. the number of edges for a Bayesian network or an image showing some representation of the instance. The set of characteristics should always include the XML-representation of the instance. The corresponding result type isStorableResult.- Specified by:
getCharacteristicsin interfaceSequenceScore- Overrides:
getCharacteristicsin classAbstractTrainableStatisticalModel- Returns:
- the characteristics of the current instance
- Throws:
Exception- if some of the characteristics could not be defined- See Also:
StorableResult
-
getNumericalCharacteristics
public NumericalResultSet getNumericalCharacteristics() throws Exception
Description copied from interface:SequenceScoreReturns the subset of numerical values that are also returned bySequenceScore.getCharacteristics().- Returns:
- the numerical characteristics of the current instance
- Throws:
Exception- if some of the characteristics could not be defined
-
getWeights
public final double[] getWeights()
This method returns a deep copy of the weights for each component.- Returns:
- the weight for each component
-
algorithmHasBeenRun
public boolean algorithmHasBeenRun()
This method indicates whether the parameters of the model has been determined by the internal algorithm.- Returns:
trueif the internal algorithm has been used to determine the parameters of the model
-
isInitialized
public boolean isInitialized()
Description copied from interface:SequenceScoreThis method can be used to determine whether the instance is initialized. If the instance is initialized you should be able to invokeSequenceScore.getLogScoreFor(Sequence).- Returns:
trueif the instance is initialized,falseotherwise
-
setAlpha
public final void setAlpha(double alpha) throws IllegalArgumentExceptionSets the parameter of the Dirichlet distribution which is used when you invoketrainto init the gammas. It is recommended to usealpha = 1(uniform distribution on a simplex).- Parameters:
alpha- the parameter of the Dirichlet distribution withalpha > 0- Throws:
IllegalArgumentException- ifalpha <= 0
-
setOutputStream
public final void setOutputStream(OutputStream o)
Sets theOutputStreamthat is used e.g. for writing information while training. It is possible to seto=null, than nothing will be written.- Parameters:
o- theOutputStream
-
getNewComponentProbs
protected void getNewComponentProbs(double[] weights) throws ExceptionEstimates the weights of each component.- Parameters:
weights- the array of weights, every element has to be non-negative and the dimension has to bedimension- Throws:
Exception- a weight is less than 0- See Also:
getNumberOfComponents()
-
setWeights
protected void setWeights(double... weights) throws IllegalArgumentExceptionSets the weights of each component.- Parameters:
weights- every element has to be non-negative, the sum of all weights has to be 1 and the dimension ofweightshas to bedimension- Throws:
IllegalArgumentException- a weight is less than 0, the sum is not equal to 1 or the dimension is incorrect- See Also:
getNumberOfComponents()
-
toXML
public StringBuffer toXML()
Description copied from interface:StorableThis method returns an XML representation asStringBufferof an instance of the implementing class.- Returns:
- the XML representation
-
getFurtherInformation
protected StringBuffer getFurtherInformation()
This method is used in the subclasses to append further information to the XML representation.- Returns:
- a part of the XML representation
- See Also:
extractFurtherInformation(StringBuffer)
-
fromXML
protected void fromXML(StringBuffer representation) throws NonParsableException
Description copied from class:AbstractTrainableStatisticalModelThis method should only be used by the constructor that works on aStringBuffer. It is the counter part ofStorable.toXML().- Specified by:
fromXMLin classAbstractTrainableStatisticalModel- Parameters:
representation- the XML representation of the model- Throws:
NonParsableException- if theStringBufferis not parsable or the representation is conflicting- See Also:
AbstractTrainableStatisticalModel.AbstractTrainableStatisticalModel(StringBuffer)
-
extractFurtherInformation
protected void extractFurtherInformation(StringBuffer xml) throws NonParsableException
This method is used in the subclasses to extract further information from the XML representation and to set these as values of the instance.- Parameters:
xml- the XML representation- Throws:
NonParsableException- if the XML representation is not parsable- See Also:
getFurtherInformation()
-
checkModelsForGibbsSampling
protected void checkModelsForGibbsSampling()
This method can be used to check whether the necessary models have implemented theSamplingComponent.
-
checkLength
protected void checkLength(int index, int l)This method checks if the lengthlof the model with indexindexis capable for the current instance. Otherwise anIllegalArgumentExceptionis thrown.- Parameters:
index- the index of the modell- the length of the model- Throws:
IllegalArgumentException- if the model instance can not be used
-
emitDataSet
public DataSet emitDataSet(int n, int... lengths) throws Exception
Description copied from interface:StatisticalModelThis method returns aDataSetobject containing artificial sequence(s).
There are two different possibilities to create a data set for a model with length 0 (homogeneous models).-
emitDataSet( int n, int l )should return a data set withnsequences of lengthl. -
emitDataSet( int n, int[] l )should return a data set withnsequences which have a sequence length corresponding to the entry in the given arrayl.
There are two different possibilities to create a data set for a model with length greater than 0 (inhomogeneous models).
emitDataSet( int n )andemitDataSet( int n, null )should return a data set withnsequences of length of the model (SequenceScore.getLength()).
The standard implementation throws anException.- Specified by:
emitDataSetin interfaceStatisticalModel- Overrides:
emitDataSetin classAbstractTrainableStatisticalModel- Parameters:
n- the number of sequences that should be contained in the returned data setlengths- the length of the sequences for a homogeneous model; for an inhomogeneous model this parameter should benullor an array of size 0.- Returns:
- a
DataSetcontaining the artificial sequence(s) - Throws:
Exception- if the emission did not succeedNotTrainedException- if the model is not trained yet- See Also:
DataSet
-
-
emitDataSetUsingCurrentParameterSet
protected abstract Sequence[] emitDataSetUsingCurrentParameterSet(int n, int... lengths) throws Exception
The method returns an array of sequences using the current parameter set.- Parameters:
n- the number of sequences to be sampledlengths- the corresponding lengths- Returns:
- an array of sequences
- Throws:
Exception- if it was impossible to sample the sequences- See Also:
StatisticalModel.emitDataSet(int, int...)
-
parseParameterSet
protected boolean parseParameterSet(int sampling, int burnInIteration) throws ExceptionThis method allows the user to parse the set of parameters with indexburnInIterationof a specificsampling(from a file).- Parameters:
sampling- the index of the samplingburnInIteration- the number of iterations that should be skipped- Returns:
trueif the parameter set could be parsed- Throws:
Exception- if something went wrong while reading or parsing the parameter set
-
parseNextParameterSet
protected boolean parseNextParameterSet() throws ExceptionThis method allows the user to parse the next set of parameters (from a file).- Returns:
trueif the parameter set could be parsed- Throws:
Exception- if something went wrong while reading or parsing the parameter set
-
initModelForSampling
protected void initModelForSampling(int starts) throws IOExceptionThis method initializes the model for the sampling. For instance this method can be used to create new files where all parameter sets will be stored.- Parameters:
starts- the number of sampling starts- Throws:
IOException- if the files could not be handled properly
-
extendSampling
protected void extendSampling(int sampling) throws ExceptionThis method prepares the model to extend an existing sampling.- Parameters:
sampling- the index of the sampling- Throws:
Exception- if the internal files could not be handled properly
-
samplingStopped
protected void samplingStopped() throws IOExceptionThis method is the opposite of the methodinitModelForSampling(int). It can be used for closing any streams of writer, ...- Throws:
IOException- if theFileWritercould not be closed properly
-
isInSamplingMode
protected boolean isInSamplingMode()
This method returnstrueif the object is currently used in a sampling, otherwisefalse.- Returns:
trueif the object is currently used in a sampling
-
finalize
protected void finalize() throws Throwable
-
draw
public static final int draw(double[] w, int start)This method draws an index of an array corresponding to the probabilities encoded in the entries of the array.- Parameters:
w- an array containing probabilities starting at positionstartstart- the start index- Returns:
- the drawn index
-
max
public static final int max(double[] w, int start, int end)This method returns the index of a maximal entry in the arraywbetween indexstartandend.- Parameters:
w- an arraystart- the start index (inclusive)end- the end index (exclusive)- Returns:
- the index of the maximal entry
-
-