SafeOutputStream.
true if for test and train dataset the sequences of
the non-reference classes have the same length as the corresponding
sequence of the reference class.
SamplingScoreBasedClassifier.burnInTest and then samples the number of
stationary steps as set in SamplingScoreBasedClassifier.params.
Sequences.Sample from a StringExtractor
using the given AlphabetContainer.
Sample from a StringExtractor
using the given AlphabetContainer and all overlapping windows of
length subsequenceLength.
Sample from a StringExtractor
using the given AlphabetContainer and a delimiter
delim.
Sample from a StringExtractor
using the given AlphabetContainer, the given delimiter
delim and all overlapping windows of length
subsequenceLength.
Sample from a given Sample and a given
length subsequenceLength.
Sample from an array of Sequences and a
given annotation.
Sample.Sample.ElementEnumerator on the given Sample
data.
enum defines different partition methods for a
Sample.Sequences that occur more
than once in one or more Samples.Sample.WeightedSampleFactory on the given
Sample(s) with Sample.WeightedSampleFactory.SortOperation sort.
Sample.WeightedSampleFactory on the given
Sample and an array of weights with
Sample.WeightedSampleFactory.SortOperation sort.
Sample.WeightedSampleFactory on the given
Sample and an array of weights with a given
length and Sample.WeightedSampleFactory.SortOperation sort.
Sample.WeightedSampleFactory on the given array of
Samples and an array of weights with a given
length and Sample.WeightedSampleFactory.SortOperation sort.
enum defines the different types of sort operations
that can be performed while creating a Sample.WeightedSampleFactory.ClassifierAssessmentAssessParameterSet that
must be used to call the method assess( ... ) of a
Sampled_RepeatedHoldOutExperiment.Sampled_RepeatedHoldOutAssessParameterSet with
empty parameter values.
Storable.
Sampled_RepeatedHoldOutAssessParameterSet with
given parameter values.
ClassifierAssessment that partitions the data
of a user-specified reference class (typically the smallest class) and
samples non-overlapping for all other classes, so that one gets the same
number of sequences (and the same lengths of the sequences) in each train and
test dataset.Sampled_RepeatedHoldOutExperiment from an array of
AbstractClassifiers and a two-dimensional array of Model
s, which are combined to additional classifiers.
Sampled_RepeatedHoldOutExperiment from a set of
AbstractClassifiers.
Sampled_RepeatedHoldOutExperiment from a set of
Models.
AbstractClassifiers and those constructed using the given
AbstractModels by a
Sampled_RepeatedHoldOutExperiment.
RecyclableSequenceEnumerator of Sequences that enumerates all k-mers that exist in a given Sample, optionally ignoring reverse complements.Sample data by extracting all k-mers.
seq using the internal parameters.
Result that contains a Sample.SampleResult from a Sample with the
annotation name and comment.
SampleResult from a Sample with the
annotation name and comment.
Storable.
SequenceIterator from the Sample
sample preserving as much annotation as possible.
LogGenDisMixFunction using the
Metropolis-Hastings algorithm.SamplingGenDisMixClassifier using the external parameters
params, a burn-in test, a set of sampling variances for the different classes,
a prior on the parameters, weights beta for the three components of the
LogGenDisMixFunction, i.e., likelihood, conditional likelihood, and prior,
and scoring functions that model the distribution for each of the classes.
SamplingGenDisMixClassifier using the external parameters
params, a burn-in test, a set of sampling variances for the different classes,
a prior on the parameters, a learning principle,
and scoring functions that model the distribution for each of the classes.
SamplingGenDisMixClassifier from its XML-representation
ParameterSet to instantiate a SamplingGenDisMixClassifier.SamplingGenDisMixClassifierParameterSet.
SamplingGenDisMixClassifierParameterSet.
SamplingGenDisMixClassifierParameterSet with a grouped sampling scheme, sampling all parameters
(and not only the free ones), and adaption of the variance.
Storable.
AbstractHMM using a sampling strategy.AbstractHMM using a sampling strategy.
Storable.
Storable.
SamplingScoringFunctions by the Metropolis-Hastings algorithm.Storable.
SamplingScoreBasedClassifier using the parameters in params,
a specified BurnInTest (or null for no burn-in test), a set of sampling variances,
which may be different for each of the classes (in analogy to equivalent sample size for the Dirichlet distribution),
and set set of SamplingScoringFunctions for each of the classes.
SamplingComponent that handles storing and loading sampled parameters values
to and from files.SamplingScoreBasedClassifier.ScoringFunctionSamplingComponent that uses temporary files
with name prefix outfilePrefix to store sampled parameters.
ParameterSet to instantiate a SamplingScoreBasedClassifier.SamplingScoreBasedClassifierParameterSet.
SamplingScoreBasedClassifierParameterSet with a grouped sampling scheme, sampling all parameters
(and not only the free ones), and adaption of the variance.
NormalizableScoringFunctions that can be used for
Metropolis-Hastings sampling in a SamplingScoreBasedClassifier.AbstractMixtureModel.initModelForSampling(int).
SamplingComponent.extendSampling(int, boolean).
Sequence seq beginning at position
start.
sequence.
Sample to a file f.
Sequences including their
SequenceAnnotations into a OutputStream.
StringBuffer representing these as XML.
Storable.
ScoringFunctions are used during the parallel computation.
ScoreBasedPerformanceMeasureDefinitions.ThresholdMeasurePair.
ScoringFunction based classifier.ScoreClassifier from a given
ScoreClassifierParameterSet and ScoringFunctions .
Storable.
Parameters for any
ScoreClassifier.ScoreClassifierParameterSet with empty parameter
values.
Storable.
ScoreClassifier.SamplingScoringFunctions
LogPrior that defines a Gaussian prior on the parameters
of a set of NormalizableScoringFunctions
and a set of class parameters.SeparateGaussianLogPrior from a set of base
variances vars, a set of class variances
classVars and a set of class means classMus.
Storable.
LogPrior that defines a Laplace prior on the parameters
of a set of NormalizableScoringFunctions
and a set of class parameters.SeparateLaplaceLogPrior from a set of base
variances vars, a set of class variances
classVars and a set of class means classMus.
Storable.
SeparateLogPrior using the class-specific base
variances vars, the variances classVars and the
means classMus for the class parameters.
Storable.
Sequence with the given AlphabetContainer
and the given annotation, but without the content.
Sequences.Sequence.CompositeSequence
for Sequences with a simple AlphabetContainer.
Sample of Sequence.CompositeSequences.
Sequence.RecursiveSequence on the Sequence
seq with the AlphabetContainer alphabet
and the annotation annotation.
Sequence.RecursiveSequence on the Sequence
seq with the AlphabetContainer alphabet
using the annotation of the given Sequence.
Sample of Sequence.SubSequences of defined length.
Sequence.SubSequence of
defined length for Sequences with a simple
AlphabetContainer.
Sequence.SequenceAnnotation of type type with
identifier identifier and additional annotation (that does
not fit the SequenceAnnotation definitions) given as a
Result result.
SequenceAnnotation of type type with
identifier identifier and additional annotation (that does
not fit the SequenceAnnotation definitions) given as an array of
Results results.
SequenceAnnotation of type type with
identifier identifier and additional annotation (that does
not fit the SequenceAnnotation definitions) given as an array of
Results additionalAnnotation.
SequenceAnnotation of type type with
identifier identifier and additional annotation (that does
not fit the SequenceAnnotation definitions) given as a
Collection of Results results.
Storable.
AbstractStringExtractor to annotate a String
which will be parsed to a Sequence.RecyclableSequenceEnumerator on user-specified Sequences.Sequences sequences.
Collection of Sequences sequences.
SequenceIterator with maximal length.
Sample from a SequenceIterator.
ParameterSet containing all parameters necessary
to construct an Object that implements
InstantiableFromParameterSet.InstanceParameterSet having empty parameter values.
SequenceScoringParameterSet having empty parameter
values.
Storable.
SequenceScoringParameterSet from an
AlphabetContainer and the length of a sequence.
SequenceScoringParameterSet for an object that can
handle sequences of variable length and with the
AlphabetContainer alphabet.
AbstractTerminationCondition.parameter.
ScoringFunctions funs.
AbstractModel.setNewAlphabetContainerInstance(AlphabetContainer) and not be
made public.
train to init the gammas.
Parameter to
defaultValue.
true (which is the default), the temporary files for storing sampled parameter
values are deleted on exit of the program.
StructureLearner.
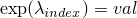 .
.
AbstractBurnInTest.
StringBuffer.
help.
helpfile.
index.
parameters.
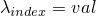 .
.
ProgressUpdater.setValue(int), so a
value of max indicates the end of the supervised method call.
ParameterSet whose validity
depends on the value of this Parameter.
AlphabetContainer
for the current classifier.
AlphabetContainer
for the current model.
i
NullProgressUpdater.setOffset() is called the current value
will be added to every value set by
NullProgressUpdater.setValue(int).
OutputStream that is used e.g. for writing information
during training.
OutputStream for the model.
OutputStream that is used e.g. for writing information
while training.
OutputStream that is used e.g. for writing information
while training.
Parameter for symbol symbol and
context context to Parameter par.
TransitionElement.offset used for several methods (cf. see tags).
TransitionElement.
params beginning at index start.
params between start and
start + ScoringFunction.getNumberOfParameters() - 1
index
ParameterSet of this
Parameter to parent.
ParameterSetContainer of this
ParameterSet to parent.
SequenceAnnotationParser that can be used to
write this SampleResult including annotations on the contained Sequences
to a file
path
data using weights.
CollectionParameter.isRangeable() to
rangeable.
SimpleParameter.isRangeable() to
rangeable.
val for the root node child.
sel depending on the new
selection b.
key to the value
of selected.
RangeParameter to one of
LIST, RANGE or NO.
String to be parsed.
SamplingScoreBasedClassifier.
train-method and sets for a
given sample the sample that should be used for train.
ParameterValidator for this SimpleParameter.
parents[0],...
- setValue(Object) -
Method in class de.jstacs.parameters.CollectionParameter
- Sets the selected value to the one that is specified by the key
value.
- setValue(Object) -
Method in class de.jstacs.parameters.EnumParameter
-
- setValue(Object) -
Method in class de.jstacs.parameters.FileParameter
-
- setValue(Object) -
Method in class de.jstacs.parameters.MultiSelectionCollectionParameter
-
- setValue(Object) -
Method in class de.jstacs.parameters.Parameter
- Sets the value of this
Parameter to value.
- setValue(Object) -
Method in class de.jstacs.parameters.ParameterSetContainer
-
- setValue(Object) -
Method in class de.jstacs.parameters.RangeParameter
-
- setValue(Object) -
Method in class de.jstacs.parameters.SimpleParameter
-
- setValue(double) -
Method in class de.jstacs.sampling.AbstractBurnInTest
-
- setValue(double) -
Method in interface de.jstacs.sampling.BurnInTest
- This method can be used to fill the internal memory with the values that
will be used to determine the length of the burn-in phase.
- setValue(double) -
Method in class de.jstacs.sampling.SimpleBurnInTest
- Deprecated.
- setValue(double) -
Method in class de.jstacs.scoringFunctions.directedGraphicalModels.Parameter
- Sets the current value of this parameter.
- setValue(int) -
Method in class de.jstacs.utils.DefaultProgressUpdater
-
- setValue(int) -
Method in class de.jstacs.utils.GUIProgressUpdater
-
- setValue(int) -
Method in class de.jstacs.utils.NullProgressUpdater
-
- setValue(int) -
Method in interface de.jstacs.utils.ProgressUpdater
- Sets the current value the supervised process has reached.
- setValue(int) -
Method in class de.jstacs.utils.TimeLimitedProgressUpdater
-
- setValueFromTag(String, Object) -
Method in class de.jstacs.parameters.ParameterSetTagger
- This method allows to easily set the value of a parameter defined by the tag.
- setValues(String) -
Method in class de.jstacs.parameters.RangeParameter
- Sets a list of values from a
String containing a space separated
list of values.
- setValues(Object, int, Object, RangeParameter.Scale) -
Method in class de.jstacs.parameters.RangeParameter
- Sets the values of this
RangeParameter as a range of values,
specified by a start value, a last value, a number of steps between these
values (without the last value) and a scale in that the values between
the first and the last value are chosen.
- setValuesInLogScale(boolean, double, Object, int, Object) -
Method in class de.jstacs.parameters.RangeParameter
- This method enables you to set a list of values in an easy manner.
- setWeight(double) -
Method in class de.jstacs.models.phylo.PhyloNode
- This method set the weight (length, rate ...) for the incoming edge
- setWeights(double...) -
Method in class de.jstacs.classifier.scoringFunctionBased.gendismix.GenDisMixClassifier
- This method set the weights for the summand of the function.
- setWeights(double...) -
Method in class de.jstacs.models.mixture.AbstractMixtureModel
- Sets the weights of each component.
- SFBasedOptimizableFunction - Class in de.jstacs.classifier.scoringFunctionBased
- This abstract class is the basis of all multi-threaded
OptimizableFunctions that are based on ScoringFunctions. - SFBasedOptimizableFunction(int, ScoringFunction[], Sample[], double[][], LogPrior, boolean, boolean) -
Constructor for class de.jstacs.classifier.scoringFunctionBased.SFBasedOptimizableFunction
- Creates an instance with the underlying infrastructure.
- shallBeRanged() -
Method in class de.jstacs.parameters.RangeParameter
- Returns one of
LIST, RANGE or NO depending on
the input used to specify this RangeParameter.
- SharedStructureClassifier - Class in de.jstacs.models.discrete.inhomogeneous.shared
- This class enables you to learn the structure on all classes of the
classifier together.
- SharedStructureClassifier(int, StructureLearner.ModelType, byte, StructureLearner.LearningType, FSDAGModel...) -
Constructor for class de.jstacs.models.discrete.inhomogeneous.shared.SharedStructureClassifier
- Creates a new
SharedStructureClassifier from given
FSDAGModels.
- SharedStructureClassifier(StringBuffer) -
Constructor for class de.jstacs.models.discrete.inhomogeneous.shared.SharedStructureClassifier
- The standard constructor for the interface
Storable.
- SharedStructureMixture - Class in de.jstacs.models.discrete.inhomogeneous.shared
- This class handles a mixture of models with the same structure that is
learned via EM.
- SharedStructureMixture(FSDAGModel[], StructureLearner.ModelType, byte, int, double, TerminationCondition) -
Constructor for class de.jstacs.models.discrete.inhomogeneous.shared.SharedStructureMixture
- Creates a new
SharedStructureMixture instance which estimates the
component probabilities/weights.
- SharedStructureMixture(FSDAGModel[], StructureLearner.ModelType, byte, int, double[], double, TerminationCondition) -
Constructor for class de.jstacs.models.discrete.inhomogeneous.shared.SharedStructureMixture
- Creates a new
SharedStructureMixture instance with fixed
component weights.
- SharedStructureMixture(FSDAGModel[], StructureLearner.ModelType, byte, int, boolean, double[], double, TerminationCondition) -
Constructor for class de.jstacs.models.discrete.inhomogeneous.shared.SharedStructureMixture
- Creates a new
SharedStructureMixture instance with all relevant
values.
- SharedStructureMixture(StringBuffer) -
Constructor for class de.jstacs.models.discrete.inhomogeneous.shared.SharedStructureMixture
- The standard constructor for the interface
Storable.
- shortcut -
Variable in class de.jstacs.classifier.scoringFunctionBased.SFBasedOptimizableFunction
- These shortcuts indicate the beginning of a new part in the parameter vector.
- ShortSequence - Class in de.jstacs.data.sequences
- This class is for sequences with the alphabet symbols encoded as
shortss and can therefore be used for discrete
AlphabetContainers with alphabets that use many different symbols. - ShortSequence(AlphabetContainer, short[]) -
Constructor for class de.jstacs.data.sequences.ShortSequence
- Creates a new
ShortSequence from an array of short-
encoded alphabet symbols.
- ShortSequence(AlphabetContainer, String) -
Constructor for class de.jstacs.data.sequences.ShortSequence
- Creates a new
ShortSequence from a String representation
using the default delimiter.
- ShortSequence(AlphabetContainer, SequenceAnnotation[], String, String) -
Constructor for class de.jstacs.data.sequences.ShortSequence
- Creates a new
ShortSequence from a String representation
using the delimiter delim.
- ShortSequence(AlphabetContainer, SequenceAnnotation[], SymbolExtractor) -
Constructor for class de.jstacs.data.sequences.ShortSequence
- Creates a new
ShortSequence from a SymbolExtractor.
- shouldBeNormalized() -
Method in class de.jstacs.classifier.scoringFunctionBased.gendismix.GenDisMixClassifierParameterSet
- This method indicates if a normalization shall be used while
optimization.
- showImage(String, BufferedImage) -
Static method in class de.jstacs.utils.REnvironment
- Enables you to show an image.
- showImage(String, BufferedImage, int) -
Static method in class de.jstacs.utils.REnvironment
- Enables you to show an image.
- SignificantMotifOccurrencesFinder - Class in de.jstacs.motifDiscovery
- This class enables the user to predict motif occurrences given a specific significance level.
- SignificantMotifOccurrencesFinder(MotifDiscoverer, SignificantMotifOccurrencesFinder.RandomSeqType, boolean, int, double) -
Constructor for class de.jstacs.motifDiscovery.SignificantMotifOccurrencesFinder
- This constructor creates an instance of
SignificantMotifOccurrencesFinder that uses the given SignificantMotifOccurrencesFinder.RandomSeqType to determine the siginificance level.
- SignificantMotifOccurrencesFinder(MotifDiscoverer, SignificantMotifOccurrencesFinder.RandomSeqType, SignificantMotifOccurrencesFinder.JoinMethod, boolean, int, double) -
Constructor for class de.jstacs.motifDiscovery.SignificantMotifOccurrencesFinder
- This constructor creates an instance of
SignificantMotifOccurrencesFinder that uses the given SignificantMotifOccurrencesFinder.RandomSeqType to determine the siginificance level.
- SignificantMotifOccurrencesFinder(MotifDiscoverer, Sample, double[], double) -
Constructor for class de.jstacs.motifDiscovery.SignificantMotifOccurrencesFinder
- This constructor creates an instance of
SignificantMotifOccurrencesFinder that uses a Sample to determine the siginificance level.
- SignificantMotifOccurrencesFinder(MotifDiscoverer, SignificantMotifOccurrencesFinder.JoinMethod, Sample, double[], double) -
Constructor for class de.jstacs.motifDiscovery.SignificantMotifOccurrencesFinder
- This constructor creates an instance of
SignificantMotifOccurrencesFinder that uses a Sample to determine the siginificance level.
- SignificantMotifOccurrencesFinder.JoinMethod - Interface in de.jstacs.motifDiscovery
- Interface for methods that combine several profiles over the same sequence
into one common profile
- SignificantMotifOccurrencesFinder.RandomSeqType - Enum in de.jstacs.motifDiscovery
-
- SignificantMotifOccurrencesFinder.SumOfProbabilities - Class in de.jstacs.motifDiscovery
- Joins several profiles containing log-probabilities into one profile containing
the logarithm of the sum of the probabilities of the single profiles.
- SignificantMotifOccurrencesFinder.SumOfProbabilities() -
Constructor for class de.jstacs.motifDiscovery.SignificantMotifOccurrencesFinder.SumOfProbabilities
-
- SilentEmission - Class in de.jstacs.models.hmm.states.emissions
- This class implements a silent emission which is used to create silent states.
- SilentEmission() -
Constructor for class de.jstacs.models.hmm.states.emissions.SilentEmission
- The main constructor.
- SilentEmission(StringBuffer) -
Constructor for class de.jstacs.models.hmm.states.emissions.SilentEmission
- The standard constructor for the interface
Storable.
- SimpleBurnInTest - Class in de.jstacs.sampling
- Deprecated.
- SimpleBurnInTest(int) -
Constructor for class de.jstacs.sampling.SimpleBurnInTest
- Deprecated. This is the main constructor that creates an instance of
SimpleBurnInTest with fixed burn-in length.
- SimpleBurnInTest(StringBuffer) -
Constructor for class de.jstacs.sampling.SimpleBurnInTest
- Deprecated. The standard constructor for the interface
Storable.
- SimpleCosts - Class in de.jstacs.algorithms.alignment.cost
- Class for simple costs with costs
mismatch for a mismatch,
costs start to start a new gap, costs elong to
elongate a gap by one position and costs of match for a match. - SimpleCosts(double, double, double, double) -
Constructor for class de.jstacs.algorithms.alignment.cost.SimpleCosts
- Creates a new instance of simple costs with costs
mismatch for a mismatch, costs start to
start a new gap, costs elong to elongate a gap by one
position and costs of match for a match.
- SimpleDifferentiableState - Class in de.jstacs.models.hmm.states
- This class implements a
State based on Emission that allows to reuse Emissions for different States. - SimpleDifferentiableState(DifferentiableEmission, String, boolean) -
Constructor for class de.jstacs.models.hmm.states.SimpleDifferentiableState
- This is the constructor of a
SimpleState.
- SimpleDiscreteSequence - Class in de.jstacs.data.sequences
- This is the main class for any discrete sequence.
- SimpleDiscreteSequence(AlphabetContainer, SequenceAnnotation[]) -
Constructor for class de.jstacs.data.sequences.SimpleDiscreteSequence
- This constructor creates a new
SimpleDiscreteSequence with the
AlphabetContainer container and the annotation
annotation but without the content.
- SimpleGaussianSumLogPrior - Class in de.jstacs.classifier.scoringFunctionBased.logPrior
- This class implements a prior that is a product of Gaussian distributions
with mean 0 and equal variance for each parameter.
- SimpleGaussianSumLogPrior(double) -
Constructor for class de.jstacs.classifier.scoringFunctionBased.logPrior.SimpleGaussianSumLogPrior
- Creates a new
SimpleGaussianSumLogPrior with mean 0 and variance
sigma for all parameters, including the class parameters.
- SimpleGaussianSumLogPrior(StringBuffer) -
Constructor for class de.jstacs.classifier.scoringFunctionBased.logPrior.SimpleGaussianSumLogPrior
- The standard constructor for the interface
Storable.
- SimpleHistory - Class in de.jstacs.motifDiscovery.history
- This class implements a simple history that has a limited memory that will be
used cyclicly.
- SimpleHistory(int) -
Constructor for class de.jstacs.motifDiscovery.history.SimpleHistory
- This constructor creates a simple history with limited memory.
- SimpleHistory(int, boolean, boolean, boolean) -
Constructor for class de.jstacs.motifDiscovery.history.SimpleHistory
- This constructor creates a simple history with limited memory.
- SimpleHistory(StringBuffer) -
Constructor for class de.jstacs.motifDiscovery.history.SimpleHistory
- This is the constructor for the interface
Storable.
- SimpleParameter - Class in de.jstacs.parameters
- Class for a "simple" parameter.
- SimpleParameter(StringBuffer) -
Constructor for class de.jstacs.parameters.SimpleParameter
- The standard constructor for the interface
Storable.
- SimpleParameter(DataType, String, String, boolean) -
Constructor for class de.jstacs.parameters.SimpleParameter
- Constructor for a
SimpleParameter without
ParameterValidator.
- SimpleParameter(DataType, String, String, boolean, Object) -
Constructor for class de.jstacs.parameters.SimpleParameter
- Constructor for a
SimpleParameter without
ParameterValidator but with a default value.
- SimpleParameter(DataType, String, String, boolean, ParameterValidator) -
Constructor for class de.jstacs.parameters.SimpleParameter
- Constructor for a
SimpleParameter with a
ParameterValidator.
- SimpleParameter(DataType, String, String, boolean, ParameterValidator, Object) -
Constructor for class de.jstacs.parameters.SimpleParameter
- Constructor for a
SimpleParameter with validator and default
value.
- SimpleParameter.DatatypeNotValidException - Exception in de.jstacs.parameters
- Class for an
Exception that can be thrown if the provided
int-value that represents a data type is not one of the
values defined in DataType. - SimpleParameter.DatatypeNotValidException(String) -
Constructor for exception de.jstacs.parameters.SimpleParameter.DatatypeNotValidException
- Creates a new
SimpleParameter.DatatypeNotValidException with an error
message.
- SimpleParameter.IllegalValueException - Exception in de.jstacs.parameters
- This exception is thrown if a parameter is not valid.
- SimpleParameter.IllegalValueException(String) -
Constructor for exception de.jstacs.parameters.SimpleParameter.IllegalValueException
- Creates a new
SimpleParameter.IllegalValueException with the reason of the
exception reason as error message.
- SimpleParameterSet - Class in de.jstacs.parameters
- Class for a
ParameterSet that is constructed from an array of Parameters
and thus does nothing in the method SimpleParameterSet.loadParameters(). - SimpleParameterSet(Parameter...) -
Constructor for class de.jstacs.parameters.SimpleParameterSet
- Creates a new
SimpleParameterSet from an array of Parameters.
- SimpleParameterSet(StringBuffer) -
Constructor for class de.jstacs.parameters.SimpleParameterSet
- The standard constructor for the interface
Storable.
- SimpleReferenceConstraint - Class in de.jstacs.parameters.validation
- Class for a
ReferenceConstraint that checks for "simple"
conditions as defined in the interface Constraint. - SimpleReferenceConstraint(SimpleParameter, int) -
Constructor for class de.jstacs.parameters.validation.SimpleReferenceConstraint
- Creates a new
SimpleReferenceConstraint from a reference
SimpleParameter and a comparison operator, which is one of the
values defined in the Constraint interface.
- SimpleReferenceConstraint(StringBuffer) -
Constructor for class de.jstacs.parameters.validation.SimpleReferenceConstraint
- The standard constructor for the interface
Storable.
- SimpleResult - Class in de.jstacs.results
- Abstract class for a
Result with a value of a primitive data type or
String. - SimpleResult(String, String, DataType) -
Constructor for class de.jstacs.results.SimpleResult
- The main constructor which takes the main information of a result.
- SimpleResult(StringBuffer) -
Constructor for class de.jstacs.results.SimpleResult
- This is the constructor for
Storable.
- SimpleSamplingState - Class in de.jstacs.models.hmm.states
- This class implements a state that can be used for a HMM that obtains its parameters from sampling.
- SimpleSamplingState(SamplingEmission, String, boolean) -
Constructor for class de.jstacs.models.hmm.states.SimpleSamplingState
- This constructor creates a state that can be used in a HMM that obtains its parameters from sampling.
- SimpleSequenceAnnotationParser - Class in de.jstacs.data.sequences.annotation
- This class implements a naive
SequenceAnnotationParser which simply paste the comments into SequenceAnnotation. - SimpleSequenceAnnotationParser() -
Constructor for class de.jstacs.data.sequences.annotation.SimpleSequenceAnnotationParser
- The constructor of a
SimpleSequenceAnnotationParser which simply paste the comments into SequenceAnnotation.
- SimpleSequenceIterator - Class in de.jstacs.data.bioJava
- Class that implements the
SequenceIterator interface of BioJava in a
simple way, backed by an array of Sequences. - SimpleSequenceIterator(Sequence...) -
Constructor for class de.jstacs.data.bioJava.SimpleSequenceIterator
- Creates a new
SimpleSequenceIterator from an array of
Sequences.
- SimpleState - Class in de.jstacs.models.hmm.states
- This class implements a
State based on Emission that allows to reuse Emissions for different States. - SimpleState(Emission, String, boolean) -
Constructor for class de.jstacs.models.hmm.states.SimpleState
- This is the constructor of a
SimpleState.
- SimpleStaticConstraint - Class in de.jstacs.parameters.validation
- Class for a
Constraint that checks values against static values using
the comparison operators defined in the interface Constraint. - SimpleStaticConstraint(Number, int) -
Constructor for class de.jstacs.parameters.validation.SimpleStaticConstraint
- Creates a new
SimpleStaticConstraint from a Number
-reference and a comparison operator as defined in Constraint.
- SimpleStaticConstraint(String, int) -
Constructor for class de.jstacs.parameters.validation.SimpleStaticConstraint
- Creates a new
SimpleStaticConstraint from a String
-reference and a comparison operator as defined in Constraint.
- SimpleStaticConstraint(StringBuffer) -
Constructor for class de.jstacs.parameters.validation.SimpleStaticConstraint
- The standard constructor for the interface
Storable.
- SimpleStringExtractor - Class in de.jstacs.io
- This is a simple class that extracts
Strings. - SimpleStringExtractor(String...) -
Constructor for class de.jstacs.io.SimpleStringExtractor
- This constructor packs the
Strings in an instance of
SimpleStringExtractor.
- simplify() -
Method in class de.jstacs.parameters.CollectionParameter
-
- simplify() -
Method in class de.jstacs.parameters.FileParameter
-
- simplify() -
Method in class de.jstacs.parameters.MultiSelectionCollectionParameter
-
- simplify() -
Method in class de.jstacs.parameters.Parameter
- Simplifies the
Parameter and its contents to the relevant
information.
- simplify() -
Method in class de.jstacs.parameters.ParameterSet
- Simplifies all
Parameters in this ParameterSet.
- simplify() -
Method in class de.jstacs.parameters.ParameterSetContainer
-
- simplify() -
Method in class de.jstacs.parameters.RangeParameter
-
- simplify() -
Method in class de.jstacs.parameters.SimpleParameter
-
- SingleHiddenMotifMixture - Class in de.jstacs.models.mixture.motif
- This class enables the user to search for a single motif in a sequence.
- SingleHiddenMotifMixture(Model, Model, boolean, int, double[], double[], PositionPrior, AbstractMixtureModel.Algorithm, double, TerminationCondition, AbstractMixtureModel.Parameterization, int, int, BurnInTest) -
Constructor for class de.jstacs.models.mixture.motif.SingleHiddenMotifMixture
- Creates a new
SingleHiddenMotifMixture.
- SingleHiddenMotifMixture(Model, Model, boolean, int, double[], PositionPrior, double, TerminationCondition, AbstractMixtureModel.Parameterization) -
Constructor for class de.jstacs.models.mixture.motif.SingleHiddenMotifMixture
- Creates a new
SingleHiddenMotifMixture using EM and estimating
the probability for finding a motif.
- SingleHiddenMotifMixture(Model, Model, boolean, int, double, PositionPrior, double, TerminationCondition, AbstractMixtureModel.Parameterization) -
Constructor for class de.jstacs.models.mixture.motif.SingleHiddenMotifMixture
- Creates a new
SingleHiddenMotifMixture using EM and fixed
probability for finding a motif.
- SingleHiddenMotifMixture(StringBuffer) -
Constructor for class de.jstacs.models.mixture.motif.SingleHiddenMotifMixture
- The standard constructor for the interface
Storable.
- SinglePositionSequenceAnnotation - Class in de.jstacs.data.sequences.annotation
- Class for some annotations that consist mainly of one position on a sequence.
- SinglePositionSequenceAnnotation(SinglePositionSequenceAnnotation.Type, String, int) -
Constructor for class de.jstacs.data.sequences.annotation.SinglePositionSequenceAnnotation
- Creates a new
SinglePositionSequenceAnnotation of type
type with identifier identifier and position
position.
- SinglePositionSequenceAnnotation(SinglePositionSequenceAnnotation.Type, String, int, Result...) -
Constructor for class de.jstacs.data.sequences.annotation.SinglePositionSequenceAnnotation
- Creates a new
SinglePositionSequenceAnnotation of type
type with identifier identifier, position
position and additional annotations
additionalAnnotation.
- SinglePositionSequenceAnnotation(StringBuffer) -
Constructor for class de.jstacs.data.sequences.annotation.SinglePositionSequenceAnnotation
- The standard constructor for the interface
Storable.
- SinglePositionSequenceAnnotation.Type - Enum in de.jstacs.data.sequences.annotation
- This
enum defines possible types of a
SinglePositionSequenceAnnotation. - SkewNormalLikeScoringFunction - Class in de.jstacs.scoringFunctions.mix.motifSearch
- This class implements a skew normal like discrete truncated distribution.
- SkewNormalLikeScoringFunction(int, int, double, double, double) -
Constructor for class de.jstacs.scoringFunctions.mix.motifSearch.SkewNormalLikeScoringFunction
- This is the main constructor if the parameters are fixed.
- SkewNormalLikeScoringFunction(int, int, boolean, double, double, boolean, double, double, boolean, double, double, int) -
Constructor for class de.jstacs.scoringFunctions.mix.motifSearch.SkewNormalLikeScoringFunction
- This is the constructor that allows the most flexible handling of the parameters.
- SkewNormalLikeScoringFunction(StringBuffer) -
Constructor for class de.jstacs.scoringFunctions.mix.motifSearch.SkewNormalLikeScoringFunction
- This is the constructor for
Storable.
- skip(int) -
Method in class de.jstacs.models.discrete.inhomogeneous.SequenceIterator
- This method skips some position.
- skipLastClassifiersDuringClassifierTraining -
Variable in class de.jstacs.classifier.assessment.ClassifierAssessment
- Skip last classifier.
- SmallDifferenceOfFunctionEvaluationsCondition - Class in de.jstacs.algorithms.optimization.termination
- This class implements a
TerminationCondition that stops an optimization
if the difference of the current and the last function evaluations will be small, i.e.,
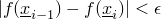 .
. - SmallDifferenceOfFunctionEvaluationsCondition(double) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallDifferenceOfFunctionEvaluationsCondition
- This constructor creates an instance that stops the optimization if the difference of the
current and the last function evaluations is smaller than
epsilon.
- SmallDifferenceOfFunctionEvaluationsCondition(SmallDifferenceOfFunctionEvaluationsCondition.SmallDifferenceOfFunctionEvaluationsConditionParameterSet) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallDifferenceOfFunctionEvaluationsCondition
- This is the main constructor creating an instance from a given parameter set.
- SmallDifferenceOfFunctionEvaluationsCondition(StringBuffer) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallDifferenceOfFunctionEvaluationsCondition
- The standard constructor for the interface
Storable.
- SmallDifferenceOfFunctionEvaluationsCondition.SmallDifferenceOfFunctionEvaluationsConditionParameterSet - Class in de.jstacs.algorithms.optimization.termination
- This class implements the parameter set for a
SmallDifferenceOfFunctionEvaluationsCondition. - SmallDifferenceOfFunctionEvaluationsCondition.SmallDifferenceOfFunctionEvaluationsConditionParameterSet() -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallDifferenceOfFunctionEvaluationsCondition.SmallDifferenceOfFunctionEvaluationsConditionParameterSet
- This constructor creates an empty parameter set.
- SmallDifferenceOfFunctionEvaluationsCondition.SmallDifferenceOfFunctionEvaluationsConditionParameterSet(StringBuffer) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallDifferenceOfFunctionEvaluationsCondition.SmallDifferenceOfFunctionEvaluationsConditionParameterSet
- The standard constructor for the interface
Storable.
- SmallDifferenceOfFunctionEvaluationsCondition.SmallDifferenceOfFunctionEvaluationsConditionParameterSet(double) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallDifferenceOfFunctionEvaluationsCondition.SmallDifferenceOfFunctionEvaluationsConditionParameterSet
- This constructor creates a filled instance of a parameters set.
- SmallGradientConditon - Class in de.jstacs.algorithms.optimization.termination
- This class implements a
TerminationCondition that allows no further iteration in an optimization if the
the gradient becomes small, i.e.,
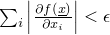 .
. - SmallGradientConditon(double) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallGradientConditon
- This constructor creates an instance that stops the optimization if the sum of the absolute
values of gradient components is smaller than
epsilon.
- SmallGradientConditon(SmallGradientConditon.SmallGradientConditonParameterSet) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallGradientConditon
- This is the main constructor creating an instance from a given parameter set.
- SmallGradientConditon(StringBuffer) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallGradientConditon
- The standard constructor for the interface
Storable.
- SmallGradientConditon.SmallGradientConditonParameterSet - Class in de.jstacs.algorithms.optimization.termination
- This class implements the parameter set for a
SmallStepCondition. - SmallGradientConditon.SmallGradientConditonParameterSet() -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallGradientConditon.SmallGradientConditonParameterSet
- This constructor creates an empty parameter set.
- SmallGradientConditon.SmallGradientConditonParameterSet(StringBuffer) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallGradientConditon.SmallGradientConditonParameterSet
- The standard constructor for the interface
Storable.
- SmallGradientConditon.SmallGradientConditonParameterSet(double) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallGradientConditon.SmallGradientConditonParameterSet
- This constructor creates a filled instance of a parameters set.
- SmallStepCondition - Class in de.jstacs.algorithms.optimization.termination
- This class implements a
TerminationCondition that allows no further iteration in an optimization if the
scalar product of the current and the last values of x will be small, i.e.,
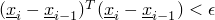 .
. - SmallStepCondition(double) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallStepCondition
- This constructor creates an instance that allows no further iteration in an optimization if the
scalar product of the current and the last values of
x is smaller than epsilon.
- SmallStepCondition(SmallStepCondition.SmallStepConditionParameterSet) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallStepCondition
- This is the main constructor creating an instance from a given parameter set.
- SmallStepCondition(StringBuffer) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallStepCondition
- The standard constructor for the interface
Storable.
- SmallStepCondition.SmallStepConditionParameterSet - Class in de.jstacs.algorithms.optimization.termination
- This class implements the parameter set for a
SmallStepCondition. - SmallStepCondition.SmallStepConditionParameterSet() -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallStepCondition.SmallStepConditionParameterSet
- This constructor creates an empty parameter set.
- SmallStepCondition.SmallStepConditionParameterSet(StringBuffer) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallStepCondition.SmallStepConditionParameterSet
- The standard constructor for the interface
Storable.
- SmallStepCondition.SmallStepConditionParameterSet(double) -
Constructor for class de.jstacs.algorithms.optimization.termination.SmallStepCondition.SmallStepConditionParameterSet
- This constructor creates a filled instance of a parameters set.
- smooth(double[]) -
Method in class de.jstacs.data.DinucleotideProperty.MeanSmoothing
-
- smooth(double[]) -
Method in class de.jstacs.data.DinucleotideProperty.MedianSmoothing
-
- smooth(double[]) -
Method in class de.jstacs.data.DinucleotideProperty.NoSmoothing
-
- smooth(double[]) -
Method in class de.jstacs.data.DinucleotideProperty.Smoothing
- Returns the smoothed version of
original.
- SoftOneOfN - Class in de.jstacs.utils.random
- This random generator returns
1-epsilon for one and equal parts
for the rest of a random vector. - SoftOneOfN(double) -
Constructor for class de.jstacs.utils.random.SoftOneOfN
- This constructor can be used for (soft) sampling one of n.
- SoftOneOfN() -
Constructor for class de.jstacs.utils.random.SoftOneOfN
- This constructor can be used for (hard) sampling one of n.
- sort(String) -
Method in class de.jstacs.results.ListResult
- This method enables you to sort the entries of this container by a
specified column.
- sort() -
Method in class de.jstacs.utils.IntList
- This method sorts the elements of the list.
- sostream -
Variable in class de.jstacs.classifier.scoringFunctionBased.ScoreClassifier
- This stream is used for comments, e.g. during the training, ... .
- sostream -
Variable in class de.jstacs.models.discrete.inhomogeneous.InhomogeneousDGM
- This stream is used for comments, computation steps/results or any other
kind of output during the training, ... etc.
- sostream -
Variable in class de.jstacs.models.hmm.AbstractHMM
- This is the stream for writing information while training.
- sostream -
Variable in class de.jstacs.models.mixture.AbstractMixtureModel
- This is the stream for writing information while training.
- source -
Variable in class de.jstacs.algorithms.graphs.Edge
- The source node.
- SparseSequence - Class in de.jstacs.data.sequences
- This class is an implementation for sequences on one alphabet with length 4.
- SparseSequence(AlphabetContainer, String) -
Constructor for class de.jstacs.data.sequences.SparseSequence
- Creates a new
SparseSequence from a String
representation.
- SparseSequence(AlphabetContainer, SymbolExtractor) -
Constructor for class de.jstacs.data.sequences.SparseSequence
- Creates a new
SparseSequence from a SymbolExtractor.
- SparseStringExtractor - Class in de.jstacs.io
- This
StringExtractor reads the Strings from a File as
the user asks for the String. - SparseStringExtractor(String) -
Constructor for class de.jstacs.io.SparseStringExtractor
- A constructor that reads the lines from a file.
- SparseStringExtractor(File) -
Constructor for class de.jstacs.io.SparseStringExtractor
- A constructor that reads the lines from a file.
- SparseStringExtractor(String, SequenceAnnotationParser) -
Constructor for class de.jstacs.io.SparseStringExtractor
- A constructor that reads the lines from a file.
- SparseStringExtractor(String, char) -
Constructor for class de.jstacs.io.SparseStringExtractor
- A constructor that reads the lines from a file and ignores
those starting with the comment character
ignore.
- SparseStringExtractor(File, char) -
Constructor for class de.jstacs.io.SparseStringExtractor
- A constructor that reads the lines from a file and ignores
those starting with the comment character
ignore.
- SparseStringExtractor(String, char, SequenceAnnotationParser) -
Constructor for class de.jstacs.io.SparseStringExtractor
- A constructor that reads the lines from a file and ignores
those starting with the comment character
ignore.
- SparseStringExtractor(String, String, SequenceAnnotationParser) -
Constructor for class de.jstacs.io.SparseStringExtractor
- A constructor that reads the lines from a file and sets the
annotation of the source to
annotation.
- SparseStringExtractor(String, char, String, SequenceAnnotationParser) -
Constructor for class de.jstacs.io.SparseStringExtractor
- A constructor that reads the lines from a file, ignores those
starting with the comment character
ignore and sets the
annotation of the source to annotation.
- SparseStringExtractor(File, char, String, SequenceAnnotationParser) -
Constructor for class de.jstacs.io.SparseStringExtractor
- A constructor that reads the lines from a file, ignores those
starting with the comment character
ignore and sets the
annotation of the source to annotation.
- spearmanCorrelation(double[], double[]) -
Static method in class de.jstacs.utils.ToolBox
- The method computes the Spearman correlation of two vectors.
- SplitSequenceAnnotationParser - Class in de.jstacs.data.sequences.annotation
- This class implements a simple
SequenceAnnotationParser which simply splits the comments by specific delimiters. - SplitSequenceAnnotationParser() -
Constructor for class de.jstacs.data.sequences.annotation.SplitSequenceAnnotationParser
- Creates a new
SplitSequenceAnnotationParser with specific delimiters, i.e., key value
delimiter "=" and annotation delimiter ";".
- SplitSequenceAnnotationParser(String, String) -
Constructor for class de.jstacs.data.sequences.annotation.SplitSequenceAnnotationParser
- Creates a new
SplitSequenceAnnotationParser with user-specified delimiters.
- standardDeviation -
Variable in class de.jstacs.models.hmm.states.emissions.continuous.PluginGaussianEmission
- Initial standard deviation.
- START_NODE -
Static variable in class de.jstacs.models.hmm.AbstractHMM
- The
String for the start node used in Graphviz annotation.
- StartDistanceForecaster - Interface in de.jstacs.algorithms.optimization
- This interface is used to determine the next start distance that will be used
in a line search.
- starts -
Variable in class de.jstacs.models.CompositeModel
- The start indices.
- starts -
Variable in class de.jstacs.models.mixture.AbstractMixtureModel
- The number of starts.
- State - Interface in de.jstacs.models.hmm
- This interface declares the methods of any state used in a hidden Markov model.
- stateList -
Variable in class de.jstacs.models.hmm.models.HigherOrderHMM
- Helper variable = only for internal use.
- states -
Variable in class de.jstacs.models.hmm.AbstractHMM
- The (hidden) states of the HMM.
- states -
Variable in class de.jstacs.models.hmm.transitions.BasicHigherOrderTransition.AbstractTransitionElement
- The states that can be visited
- StationaryDistribution - Class in de.jstacs.models.utils
- This class can be used to determine the stationary distribution.
- StationaryDistribution() -
Constructor for class de.jstacs.models.utils.StationaryDistribution
-
- stationaryIteration -
Variable in class de.jstacs.models.mixture.AbstractMixtureModel
- The number of (stationary) iterations of the Gibbs Sampler.
- statistic -
Variable in class de.jstacs.models.hmm.states.emissions.discrete.AbstractConditionalDiscreteEmission
- The array for storing the statistics for
each parameter
- statistic -
Variable in class de.jstacs.models.hmm.transitions.BasicHigherOrderTransition.AbstractTransitionElement
- The sufficient statistic for determining the parameters during sampling, viterbi or Baum-Welch training.
- StatisticalTest - Class in de.jstacs.models.utils
- This class enables the user to compute some divergences.
- StatisticalTest() -
Constructor for class de.jstacs.models.utils.StatisticalTest
-
- statisticsTransitionProb -
Variable in class de.jstacs.models.hmm.transitions.elements.DistanceBasedScaledTransitionElement
- Represents the summarized epsilons required for estimating the transition probabilities from the
context.
- statisticsTransitionProb -
Variable in class de.jstacs.models.hmm.transitions.elements.ReferenceBasedTransitionElement
- Represents the gammas required for estimating the transition probabilities not including pseudocounts.
- STEEPEST_DESCENT -
Static variable in class de.jstacs.algorithms.optimization.Optimizer
- This constant can be used to specify that the steepest descent should be
used in the
optimize-method.
- steepestDescent(DifferentiableFunction, double[], TerminationCondition, double, StartDistanceForecaster, OutputStream, Time) -
Static method in class de.jstacs.algorithms.optimization.Optimizer
- The steepest descent.
- stopThreads() -
Method in class de.jstacs.classifier.scoringFunctionBased.AbstractMultiThreadedOptimizableFunction
- This method can and should be used to stop all threads if they are not needed any longer.
- Storable - Interface in de.jstacs
- This is the root interface for all immutable objects that must be stored in
e.g. a file or a database.
- StorableResult - Class in de.jstacs.results
- Class for
Results that are Storables. - StorableResult(String, String, Storable) -
Constructor for class de.jstacs.results.StorableResult
- Creates a result for an XML representation of an object.
- StorableResult(StringBuffer) -
Constructor for class de.jstacs.results.StorableResult
- The standard constructor for the interface
Storable.
- StorableValidator - Class in de.jstacs.parameters.validation
- Class for a validator that validates instances and XML representations for
the correct class types (e.g.
- StorableValidator(Class<? extends Storable>, boolean) -
Constructor for class de.jstacs.parameters.validation.StorableValidator
- Creates a new
StorableValidator for a subclass of
AbstractModel or AbstractClassifier.
- StorableValidator(Class<? extends Storable>) -
Constructor for class de.jstacs.parameters.validation.StorableValidator
- Creates a new
StorableValidator for a subclass of
Storable.
- StorableValidator(StringBuffer) -
Constructor for class de.jstacs.parameters.validation.StorableValidator
- The standard constructor for the interface
Storable.
- StrandedLocatedSequenceAnnotationWithLength - Class in de.jstacs.data.sequences.annotation
- Class for a
SequenceAnnotation that has a position, a length and an
orientation on the strand of a Sequence. - StrandedLocatedSequenceAnnotationWithLength(int, int, StrandedLocatedSequenceAnnotationWithLength.Strand, String, String, Result...) -
Constructor for class de.jstacs.data.sequences.annotation.StrandedLocatedSequenceAnnotationWithLength
- Creates a new
StrandedLocatedSequenceAnnotationWithLength of type
type with identifier identifier and additional
annotation (that does not fit the SequenceAnnotation definitions)
given as an array of Results results.
- StrandedLocatedSequenceAnnotationWithLength(int, int, StrandedLocatedSequenceAnnotationWithLength.Strand, String, String, Collection<Result>) -
Constructor for class de.jstacs.data.sequences.annotation.StrandedLocatedSequenceAnnotationWithLength
- Creates a new
StrandedLocatedSequenceAnnotationWithLength of type
type with identifier identifier and additional
annotation (that does not fit the SequenceAnnotation definitions)
given as a Collection of Results results.
- StrandedLocatedSequenceAnnotationWithLength(int, int, StrandedLocatedSequenceAnnotationWithLength.Strand, String, String, SequenceAnnotation[], Result...) -
Constructor for class de.jstacs.data.sequences.annotation.StrandedLocatedSequenceAnnotationWithLength
- Creates a new
StrandedLocatedSequenceAnnotationWithLength of type
type with identifier identifier, additional
annotation (that does not fit the SequenceAnnotation definitions)
given as an array of Results additionalAnnotations
and sub-annotations annotations.
- StrandedLocatedSequenceAnnotationWithLength(String, String, StrandedLocatedSequenceAnnotationWithLength.Strand, LocatedSequenceAnnotation[], Result...) -
Constructor for class de.jstacs.data.sequences.annotation.StrandedLocatedSequenceAnnotationWithLength
- Creates a new
StrandedLocatedSequenceAnnotationWithLength of type
type with identifier identifier, additional
annotation (that does not fit the SequenceAnnotation definitions)
given as an array of Results additionalAnnotations
and sub-annotations annotations.
- StrandedLocatedSequenceAnnotationWithLength(StringBuffer) -
Constructor for class de.jstacs.data.sequences.annotation.StrandedLocatedSequenceAnnotationWithLength
- The standard constructor for the interface
Storable.
- StrandedLocatedSequenceAnnotationWithLength.Strand - Enum in de.jstacs.data.sequences.annotation
- This enum defines possible orientations on the strands.
- strandedness() -
Method in enum de.jstacs.data.sequences.annotation.StrandedLocatedSequenceAnnotationWithLength.Strand
- Returns the strandedness, i.e. the orientation on the strand of the
sequence as a
String.
- StrandModel - Class in de.jstacs.models.mixture
- This model handles sequences that can either lie on the forward strand or on
the reverse complementary strand.
- StrandModel(Model, int, boolean, double[], double, AbstractMixtureModel.Algorithm, double, TerminationCondition, AbstractMixtureModel.Parameterization, int, int, BurnInTest) -
Constructor for class de.jstacs.models.mixture.StrandModel
- Creates a new
StrandModel.
- StrandModel(Model, int, double[], double, TerminationCondition, AbstractMixtureModel.Parameterization) -
Constructor for class de.jstacs.models.mixture.StrandModel
- Creates an instance using EM and estimating the component probabilities.
- StrandModel(Model, int, double, double, TerminationCondition, AbstractMixtureModel.Parameterization) -
Constructor for class de.jstacs.models.mixture.StrandModel
- Creates an instance using EM and fixed component probabilities.
- StrandModel(Model, int, double[], int, int, BurnInTest) -
Constructor for class de.jstacs.models.mixture.StrandModel
- Creates an instance using Gibbs Sampling and sampling the component
probabilities.
- StrandModel(Model, int, double, int, int, BurnInTest) -
Constructor for class de.jstacs.models.mixture.StrandModel
- Creates an instance using Gibbs Sampling and fixed component
probabilities.
- StrandModel(StringBuffer) -
Constructor for class de.jstacs.models.mixture.StrandModel
- The constructor for the interface
Storable.
- StrandScoringFunction - Class in de.jstacs.scoringFunctions.mix
- This class enables the user to search on both strand.
- StrandScoringFunction(NormalizableScoringFunction, double, int, boolean, StrandScoringFunction.InitMethod) -
Constructor for class de.jstacs.scoringFunctions.mix.StrandScoringFunction
- This constructor creates a StrandScoringFunction that optimizes the usage of each strand.
- StrandScoringFunction(NormalizableScoringFunction, int, boolean, StrandScoringFunction.InitMethod, double) -
Constructor for class de.jstacs.scoringFunctions.mix.StrandScoringFunction
- This constructor creates a StrandScoringFunction that has a fixed frequency for the strand usage.
- StrandScoringFunction(StringBuffer) -
Constructor for class de.jstacs.scoringFunctions.mix.StrandScoringFunction
- This is the constructor for
Storable.
- StrandScoringFunction.InitMethod - Enum in de.jstacs.scoringFunctions.mix
- This enum defines the different types of plug-in initialization of a
StrandScoringFunction. - StringAlignment - Class in de.jstacs.algorithms.alignment
- Class for the representation of an alignment of
Strings. - StringAlignment(double, String...) -
Constructor for class de.jstacs.algorithms.alignment.StringAlignment
- This constructor creates an instance storing the aligned Strings and the costs of the alignment.
- StringExtractor - Class in de.jstacs.io
- This class implements the reader that extracts
Strings from either a
File or a String. - StringExtractor(File, int) -
Constructor for class de.jstacs.io.StringExtractor
- A constructor that reads the lines from
file.
- StringExtractor(File, int, char) -
Constructor for class de.jstacs.io.StringExtractor
- A constructor that reads the lines from
file and ignores
those starting with the comment character ignore.
- StringExtractor(File, int, String) -
Constructor for class de.jstacs.io.StringExtractor
- A constructor that reads the lines from
file and sets the
annotation of the source to annotation.
- StringExtractor(File, int, char, String) -
Constructor for class de.jstacs.io.StringExtractor
- A constructor that reads the lines from
file, ignores those
starting with the comment character ignore and sets the
annotation of the source to annotation.
- StringExtractor(String, int, String) -
Constructor for class de.jstacs.io.StringExtractor
- A constructor that reads the lines from a
String
content and sets the annotation of the source to
annotation.
- StringExtractor(String, int, char, String) -
Constructor for class de.jstacs.io.StringExtractor
- A constructor that reads the lines from a
String
content, ignores those starting with the comment character
ignore and sets the annotation of the source to
annotation.
- StructureLearner - Class in de.jstacs.models.discrete.inhomogeneous
- This class can be used to learn the structure of any discrete model.
- StructureLearner(AlphabetContainer, int, double) -
Constructor for class de.jstacs.models.discrete.inhomogeneous.StructureLearner
- Creates a new
StructureLearner for a given
AlphabetContainer, a given length and a given equivalent
sample size (ess).
- StructureLearner(AlphabetContainer, int) -
Constructor for class de.jstacs.models.discrete.inhomogeneous.StructureLearner
- Creates a
StructureLearner with equivalent sample
size (ess) = 0.
- StructureLearner.LearningType - Enum in de.jstacs.models.discrete.inhomogeneous
- This
enum defines the different types of learning that are
possible with the StructureLearner. - StructureLearner.ModelType - Enum in de.jstacs.models.discrete.inhomogeneous
- This
enum defines the different types of models that can be
learned with the StructureLearner. - structureMeasure -
Variable in class de.jstacs.scoringFunctions.directedGraphicalModels.BayesianNetworkScoringFunction
Measure that defines the network structure.
- SubclassFinder - Class in de.jstacs.utils
- Utility-class with static methods to
find all sub-classes of a certain class (or interface) within the scope
of the current class-loader
find all sub-classes of a certain class (or interface) within the scope
of the current class-loader that can be instantiated, i.e. that are neither
interfaces nor abstract
filter a set of classes by inheritance from a super-class
obtain the class of an
InstanceParameterSet that can be used to
instantiate a sub-class of InstantiableFromParameterSet. - SubclassFinder() -
Constructor for class de.jstacs.utils.SubclassFinder
-
- subSampling(int) -
Method in class de.jstacs.data.Sample
- Randomly samples elements, i.e.
- SubTensor - Class in de.jstacs.algorithms.graphs.tensor
- This Tensor can be used to extract or use only a part of a complete
Tensor. - SubTensor(Tensor, int, int) -
Constructor for class de.jstacs.algorithms.graphs.tensor.SubTensor
- This constructor creates a
SubTensor using the Tensor t for the nodes offset, offset+1, ..., offset+length-1.
- sum -
Variable in class de.jstacs.classifier.scoringFunctionBased.AbstractOptimizableFunction
- The sums of the weighted data per class and additional the total weight
sum.
- sum(double[]) -
Static method in class de.jstacs.scoringFunctions.directedGraphicalModels.structureLearning.measures.Measure
- Computes the sum of all elements in the array
ar.
- sum(double...) -
Static method in class de.jstacs.utils.ToolBox
- Computes the sum of the values in
array
- sum(int, int, double[]) -
Static method in class de.jstacs.utils.ToolBox
- Computes the sum of the values in
array starting at
start until end.
- sumNormalisation(double[]) -
Static method in class de.jstacs.utils.Normalisation
- The method does a sum-normalisation on
d and returns the the
sum of the values.
- sumNormalisation(double[], double[], int) -
Static method in class de.jstacs.utils.Normalisation
- The method does a sum-normalisation on
d.
- swap() -
Method in class de.jstacs.models.mixture.AbstractMixtureModel
- This method swaps the current component models with the alternative
model.
- symbol -
Variable in class de.jstacs.scoringFunctions.directedGraphicalModels.Parameter
- The symbol (out of some
Alphabet) this parameter
is responsible for.
- SymbolExtractor - Class in de.jstacs.io
- This class enables you to extract elements (symbols) from a given
String similar to a StringTokenizer. - SymbolExtractor(String) -
Constructor for class de.jstacs.io.SymbolExtractor
- Creates a new
SymbolExtractor using delim as
delimiter.
- SymbolExtractor(String, String) -
Constructor for class de.jstacs.io.SymbolExtractor
- Creates a new
SymbolExtractor using delim as
delimiter and string as the String to be parsed.
- SymmetricTensor - Class in de.jstacs.algorithms.graphs.tensor
- This class can be used for
Tensors with a special symmetry property. - SymmetricTensor(int, byte) -
Constructor for class de.jstacs.algorithms.graphs.tensor.SymmetricTensor
- This constructor creates an empty symmetric tensor with given dimension.
- SymmetricTensor(SymmetricTensor[], double[]) -
Constructor for class de.jstacs.algorithms.graphs.tensor.SymmetricTensor
- The constructor can be used creating a new
SymmetricTensor as
weighted sum of SymmetricTensors.
- SymmetricTensor(AsymmetricTensor) -
Constructor for class de.jstacs.algorithms.graphs.tensor.SymmetricTensor
- This constructor creates and checks a filled asymmetric tensor from an
AsymmetricTensor instance.
- SymmetricTensor(double[][][], int, byte) -
Constructor for class de.jstacs.algorithms.graphs.tensor.SymmetricTensor
- This constructor creates and checks a filled asymmetric tensor with given
dimension.